The breadth of antiviral activity of Apobec3DE in chimpanzees has been driven by positive selection
- PMID: 21835794
- PMCID: PMC3194980
- DOI: 10.1128/JVI.05046-11
The breadth of antiviral activity of Apobec3DE in chimpanzees has been driven by positive selection
Abstract
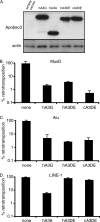
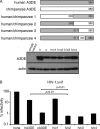
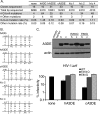
The Apobec3 family of cytidine deaminases can inhibit the replication of retroviruses and retrotransposons. Human and chimpanzee genomes encode seven Apobec3 paralogs; of these, Apobec3DE has the greatest sequence divergence between humans and chimpanzees. Here we show that even though human and chimpanzee Apobec3DEs are very divergent, the two orthologs similarly restrict long terminal repeat (LTR) and non-LTR retrotransposons (MusD and Alu, respectively). However, chimpanzee Apobec3DE also potently restricts two lentiviruses, human immunodeficiency virus type 1 (HIV-1) and the simian immunodeficiency virus (SIV) that infects African green monkeys (SIVagmTAN), unlike human Apobec3DE, which has poor antiviral activity against these same viruses. This difference between human and chimpanzee Apobec3DE in the ability to restrict retroviruses is not due to different levels of Apobec3DE protein incorporation into virions but rather to the ability of Apobec3DE to deaminate the viral genome in target cells. We further show that Apobec3DE rapidly evolved in chimpanzee ancestors approximately 2 to 6 million years ago and that this evolution drove the increased breadth of chimpanzee Apobec3DE antiviral activity to its current high activity against some lentiviruses. Despite a difference in target specificities between human and chimpanzee Apobec3DE, Apobec3DE is likely to currently play a role in host defense against retroelements in both species.
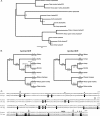
Figures
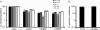


Similar articles
-
Identification of APOBEC3DE as another antiretroviral factor from the human APOBEC family.J Virol. 2006 Nov;80(21):10522-33. doi: 10.1128/JVI.01123-06. Epub 2006 Aug 18. J Virol. 2006. PMID: 16920826 Free PMC article.
-
Sole copy of Z2-type human cytidine deaminase APOBEC3H has inhibitory activity against retrotransposons and HIV-1.FASEB J. 2009 Jan;23(1):279-87. doi: 10.1096/fj.07-088781. Epub 2008 Sep 30. FASEB J. 2009. PMID: 18827027 Free PMC article.
-
The Role of the Antiviral APOBEC3 Gene Family in Protecting Chimpanzees against Lentiviruses from Monkeys.PLoS Pathog. 2015 Sep 22;11(9):e1005149. doi: 10.1371/journal.ppat.1005149. eCollection 2015 Sep. PLoS Pathog. 2015. PMID: 26394054 Free PMC article.
-
Cytidine deaminases as a weapon against retroviruses and a new target for antiviral therapy.Mini Rev Med Chem. 2008 Mar;8(3):231-8. doi: 10.2174/138955708783744047. Mini Rev Med Chem. 2008. PMID: 18336343 Review.
-
Multiple APOBEC3 restriction factors for HIV-1 and one Vif to rule them all.J Mol Biol. 2014 Mar 20;426(6):1220-45. doi: 10.1016/j.jmb.2013.10.033. Epub 2013 Nov 2. J Mol Biol. 2014. PMID: 24189052 Free PMC article. Review.
Cited by
-
APOBEC3G restricts HIV-1 to a greater extent than APOBEC3F and APOBEC3DE in human primary CD4+ T cells and macrophages.J Virol. 2013 Jan;87(1):444-53. doi: 10.1128/JVI.00676-12. Epub 2012 Oct 24. J Virol. 2013. PMID: 23097438 Free PMC article.
-
Molecular origins of APOBEC-associated mutations in cancer.DNA Repair (Amst). 2020 Oct;94:102905. doi: 10.1016/j.dnarep.2020.102905. Epub 2020 Jul 6. DNA Repair (Amst). 2020. PMID: 32818816 Free PMC article. Review.
-
Host restriction factors in retroviral infection: promises in virus-host interaction.Retrovirology. 2012 Dec 20;9:112. doi: 10.1186/1742-4690-9-112. Retrovirology. 2012. PMID: 23254112 Free PMC article. Review.
-
Molecular reconstruction of extinct LINE-1 elements and their interaction with nonautonomous elements.Mol Biol Evol. 2013 Jan;30(1):88-99. doi: 10.1093/molbev/mss202. Epub 2012 Aug 23. Mol Biol Evol. 2013. PMID: 22918960 Free PMC article.
-
The HIV-1 pandemic: does the selective sweep in chimpanzees mirror humankind's future?Retrovirology. 2013 May 24;10:53. doi: 10.1186/1742-4690-10-53. Retrovirology. 2013. PMID: 23705941 Free PMC article. Review.
References
-
- Aguiar R. S., Lovsin N., Tanuri A., Peterlin B. M. 2008. Vpr.A3A chimera inhibits HIV replication. J. Biol. Chem. 283:2518–2525 - PubMed
-
- Bartz S. R., Vodicka M. A. 1997. Production of high-titer human immunodeficiency virus type 1 pseudotyped with vesicular stomatitis virus glycoprotein. Methods 12:337–342 - PubMed
-
- Batzer M. A., Deininger P. L. 2002. Alu repeats and human genomic diversity. Nat. Rev. Genet. 3:370–379 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources

