The elusive middle domain of Hsp104 and ClpB: location and function
- PMID: 21843558
- PMCID: PMC3219823
- DOI: 10.1016/j.bbamcr.2011.07.014
The elusive middle domain of Hsp104 and ClpB: location and function
Abstract
Hsp104 in yeast and ClpB in bacteria are homologous, hexameric AAA+ proteins and Hsp100 chaperones, which function in the stress response as ring-translocases that drive protein disaggregation and reactivation. Both Hsp104 and ClpB contain a distinctive coiled-coil middle domain (MD) inserted in the first AAA+ domain, which distinguishes them from other AAA+ proteins and Hsp100 family members. Here, we focus on recent developments concerning the location and function of the MD in these hexameric molecular machines, which remains an outstanding question. While the atomic structure of the hexameric assembly of Hsp104 and ClpB remains uncertain, recent advances have illuminated that the MD is critical for the intrinsic disaggregase activity of the hexamer and mediates key functional interactions with the Hsp70 chaperone system (Hsp70 and Hsp40) that empower protein disaggregation.
Copyright © 2011 Elsevier B.V. All rights reserved.
Figures


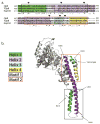
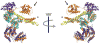

References
-
- Schirmer EC, Glover JR, Singer MA, Lindquist S. HSP100/Clp proteins: a common mechanism explains diverse functions. Trends in Biochemical Sciences. 1996;21:289–296. - PubMed
-
- Doyle SM, Wickner S. Hsp104 and ClpB: protein disaggregating machines. Trends Biochem Sci. 2009;34:40–48. - PubMed
-
- Glover JR, Lum R. Remodeling of protein aggregates by Hsp104. Protein Pept Lett. 2009;16:587–597. - PubMed
-
- Barends TR, Werbeck ND, Reinstein J. Disaggregases in 4 dimensions. Curr Opin Struct Biol. 2010;20:46–53. - PubMed
-
- Haslberger T, Bukau B, Mogk A. Towards a unifying mechanism for ClpB/Hsp104-mediated protein disaggregation and prion propagation. Biochem Cell Biol. 2010;88:63–75. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

