H3K4 tri-methylation provides an epigenetic signature of active enhancers
- PMID: 21847099
- PMCID: PMC3199384
- DOI: 10.1038/emboj.2011.295
H3K4 tri-methylation provides an epigenetic signature of active enhancers
Abstract
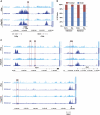
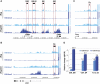
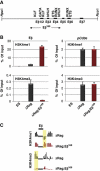
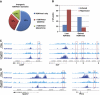
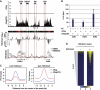
Combinations of post-translational histone modifications shape the chromatin landscape during cell development in eukaryotes. However, little is known about the modifications exactly delineating functionally engaged regulatory elements. For example, although histone H3 lysine 4 mono-methylation (H3K4me1) indicates the presence of transcriptional gene enhancers, it does not provide clearcut information about their actual position and stage-specific activity. Histone marks were, therefore, studied here at genomic loci differentially expressed in early stages of T-lymphocyte development. The concomitant presence of the three H3K4 methylation states (H3K4me1/2/3) was found to clearly reflect the activity of bona fide T-cell gene enhancers. Globally, gain or loss of H3K4me2/3 at distal genomic regions correlated with, respectively, the induction or the repression of associated genes during T-cell development. In the Tcrb gene enhancer, the H3K4me3-to-H3K4me1 ratio decreases with the enhancer's strength. Lastly, enhancer association of RNA-polymerase II (Pol II) correlated with the presence of H3K4me3 and Pol II accumulation resulted in local increase of H3K4me3. Our results suggest the existence of functional links between Pol II occupancy, H3K4me3 enrichment and enhancer activity.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures
References
-
- Adlam M, Siu G (2003) Hierarchical interactions control CD4 gene expression during thymocyte development. Immunity 18: 173–184 - PubMed
-
- Anderson MK (2006) At the crossroads: diverse roles of early thymocyte transcriptional regulators. Immunol Rev 209: 191–211 - PubMed
-
- Barski A, Cuddapah S, Cui K, Roh TY, Schones DE, Wang Z, Wei G, Chepelev I, Zhao K (2007) High-resolution profiling of histone methylations in the human genome. Cell 129: 823–837 - PubMed
-
- Bernstein BE, Mikkelsen TS, Xie X, Kamal M, Huebert DJ, Cuff J, Fry B, Meissner A, Wernig M, Plath K, Jaenisch R, Wagschal A, Feil R, Schreiber SL, Lander ES (2006) A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell 125: 315–326 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

