MicroRNA-132 targets HB-EGF upon IgE-mediated activation in murine and human mast cells
- PMID: 21853268
- PMCID: PMC11114963
- DOI: 10.1007/s00018-011-0786-3
MicroRNA-132 targets HB-EGF upon IgE-mediated activation in murine and human mast cells
Abstract
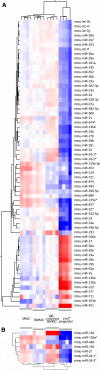
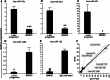
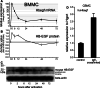
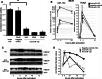
MicroRNAs provide an additional layer in the regulation of gene expression acting as repressors with several targets at the posttranscriptional level. This study describes microRNA expression patterns during differentiation and activation of mast cells. The expression levels of 567 different mouse miRNAs were compared by microarray between c-Kit+ committed progenitors, mucosal mast cells, resting and IgE-crosslinked BMMCs in vitro. The strongest upregulation of miR-132 upon IgE-mediated activation was validated in human cord blood-derived mast cells as well. HB-EGF growth factor also upregulated upon activation and was ranked high by more prediction algorithms. Co-transfection of miR-132 mimicking precursor and the 3'UTR of human Hbegf-containing luciferase vector proves that the predicted binding site is functional. In line with this, neutralization of miR-132 by anti-miR inhibitor leads to sustained production of HB-EGF protein in activated mast cells. Our data provide a novel example for negative regulation of a growth factor by an upregulated miRNA.
© Springer Basel AG 2011
Figures



Similar articles
-
Heparin-binding EGF-like growth factor and miR-1192 exert opposite effect on Runx2-induced osteogenic differentiation.Cell Death Dis. 2013 Oct 17;4(10):e868. doi: 10.1038/cddis.2013.363. Cell Death Dis. 2013. PMID: 24136232 Free PMC article.
-
IgE production in CD40/CD40L cross-talk of B and mast cells and mediator release via TGase 2 in mouse allergic asthma.Cell Signal. 2013 Jun;25(6):1514-25. doi: 10.1016/j.cellsig.2013.03.010. Epub 2013 Mar 22. Cell Signal. 2013. PMID: 23524335
-
Characterization of microRNA profile in IgE-mediated mouse BMMCs degranulation.J Microbiol Immunol Infect. 2020 Aug;53(4):550-560. doi: 10.1016/j.jmii.2018.10.006. Epub 2018 Nov 9. J Microbiol Immunol Infect. 2020. PMID: 30473142
-
Altered MicroRNA Expression Profiles in Activated Mast Cells Following IgE-FcεRI Cross-Linking with Antigen.Cell Physiol Biochem. 2015;35(6):2098-110. doi: 10.1159/000374016. Epub 2015 Apr 7. Cell Physiol Biochem. 2015. PMID: 25895812
-
CD1d expressed in mast cell surface enhances IgE production in B cells by up-regulating CD40L expression and mediator release in allergic asthma in mice.Cell Signal. 2014 May;26(5):1105-17. doi: 10.1016/j.cellsig.2014.01.029. Epub 2014 Feb 5. Cell Signal. 2014. PMID: 24509414
Cited by
-
MicroRNAs in pancreatic cancer metabolism.Nat Rev Gastroenterol Hepatol. 2012 Apr 17;9(6):334-44. doi: 10.1038/nrgastro.2012.63. Nat Rev Gastroenterol Hepatol. 2012. PMID: 22508159 Free PMC article. Review.
-
MicroRNA Involvement in Allergic and Non-Allergic Mast Cell Activation.Int J Mol Sci. 2019 Apr 30;20(9):2145. doi: 10.3390/ijms20092145. Int J Mol Sci. 2019. PMID: 31052286 Free PMC article. Review.
-
Interleukin-33 mediated regulation of microRNAs in human cord blood-derived mast cells: Implications for infection, immunity, and inflammation.PLoS One. 2024 Nov 26;19(11):e0314446. doi: 10.1371/journal.pone.0314446. eCollection 2024. PLoS One. 2024. PMID: 39591475 Free PMC article.
-
Capturing microRNA targets using an RNA-induced silencing complex (RISC)-trap approach.Proc Natl Acad Sci U S A. 2012 Dec 11;109(50):20473-8. doi: 10.1073/pnas.1218887109. Epub 2012 Nov 26. Proc Natl Acad Sci U S A. 2012. PMID: 23184980 Free PMC article.
-
MicroRNA-132 enhances transition from inflammation to proliferation during wound healing.J Clin Invest. 2015 Aug 3;125(8):3008-26. doi: 10.1172/JCI79052. Epub 2015 Jun 29. J Clin Invest. 2015. PMID: 26121747 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

