Mechanism of ubiquitylation by dimeric RING ligase RNF4
- PMID: 21857666
- PMCID: PMC3326525
- DOI: 10.1038/nsmb.2108
Mechanism of ubiquitylation by dimeric RING ligase RNF4
Abstract
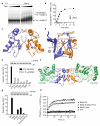
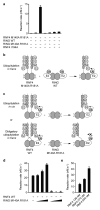
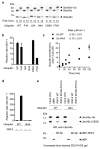
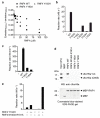
Mammalian RNF4 is a dimeric RING ubiquitin E3 ligase that ubiquitylates poly-SUMOylated proteins. We found that RNF4 bound ubiquitin-charged UbcH5a tightly but free UbcH5a weakly. To provide insight into the mechanism of RING-mediated ubiquitylation, we docked the UbcH5~ubiquitin thioester onto the RNF4 RING structure. This revealed that with E2 bound to one monomer of RNF4, the thioester-linked ubiquitin could reach across the dimer to engage the other monomer. In this model, the 'Ile44 hydrophobic patch' of ubiquitin is predicted to engage a conserved tyrosine located at the dimer interface of the RING, and mutation of these residues blocked ubiquitylation activity. Thus, dimeric RING ligases are not simply inert scaffolds that bring substrate and E2-loaded ubiquitin into close proximity. Instead, they facilitate ubiquitin transfer by preferentially binding the E2~ubiquitin thioester across the dimer and activating the thioester bond for catalysis.
Figures


References
-
- Dye BT, Schulman BA. Structural mechanisms underlying posttranslational modification by ubiquitin-like proteins. Annu. Rev. Biophys. Biomol. Struct. 2007;36:131–150. - PubMed
-
- Pickart CM, Eddins MJ. Ubiquitin: structures, functions, mechanisms. Biochim. Biophys. Acta. 2004;1695:55–72. - PubMed
-
- Deshaies RJ, Joazeiro CA. RING domain E3 ubiquitin ligases. Annu. Rev. Biochem. 2009;78:399–434. - PubMed
-
- Hay RT. SUMO: a history of modification. Mol. Cell. 2005;18:1–12. - PubMed
-
- Perry JJ, Tainer JA, Boddy MN. A SIM-ultaneous role for SUMO and ubiquitin. Trends Biochem. Sci. 2008;33:201–208. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

