The hepatitis C virus E1 glycoprotein undergoes productive folding but accelerated degradation when expressed as an individual subunit in CHO cells
- PMID: 21858229
- PMCID: PMC3157478
- DOI: 10.1371/journal.pone.0023838
The hepatitis C virus E1 glycoprotein undergoes productive folding but accelerated degradation when expressed as an individual subunit in CHO cells
Abstract
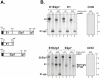
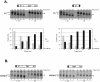
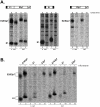
Hepatitis C Virus E1E2 heterodimers are components of the viral spike. Although there is a general agreement on the necessity of the co-expression of both E1 and E2 on a single coding unit for their productive folding and assembly, in a previous study using an in vitro system we obtained strong indications that E1 can achieve folding in absence of E2. Here, we have studied the folding pathway of unescorted E1 from stably expressing CHO cells, compared to the folding observed in presence of the E2 protein. A DTT-resistant conformation is achieved by E1 in both situations, consistent with the presence of an E2-independent oxidative pathway. However, while the E1E2 heterodimer is stable inside cells, E1 expressed alone is degraded within a few hours. On the other hand, the oxidation and stability of individually expressed E2 subunits is dependent on E1 co-expression. These data are consistent with E1 and E2 assisting each other for correct folding via different mechanisms: E2 assists E1 by stabilizing a semi-native conformation meanwhile E1 drives E2 towards a productive folding pathway.
Conflict of interest statement
Figures
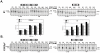
Similar articles
-
Folding and dimerization of hepatitis C virus E1 and E2 glycoproteins in stably transfected CHO cells.Virology. 2005 Feb 5;332(1):438-53. doi: 10.1016/j.virol.2004.11.034. Virology. 2005. PMID: 15661174
-
The transmembrane domain of the hepatitis C virus E2 glycoprotein is required for correct folding of the E1 glycoprotein and native complex formation.Virology. 2001 Jan 5;279(1):58-68. doi: 10.1006/viro.2000.0693. Virology. 2001. PMID: 11145889
-
Folding of hepatitis C virus E1 glycoprotein in a cell-free system.J Virol. 2001 Nov;75(22):11205-17. doi: 10.1128/JVI.75.22.11205-11217.2001. J Virol. 2001. PMID: 11602760 Free PMC article.
-
Computational Modeling of Hepatitis C Virus Envelope Glycoprotein Structure and Recognition.Front Immunol. 2018 May 28;9:1117. doi: 10.3389/fimmu.2018.01117. eCollection 2018. Front Immunol. 2018. PMID: 29892287 Free PMC article. Review.
-
Structure and Function of the Hepatitis C Virus Envelope Glycoproteins E1 and E2: Antiviral and Vaccine Targets.ACS Infect Dis. 2016 Nov 11;2(11):749-762. doi: 10.1021/acsinfecdis.6b00110. Epub 2016 Aug 16. ACS Infect Dis. 2016. PMID: 27933781 Review.
Cited by
-
Hepatitis C virus E1 recruits high-density lipoprotein to support infectivity and evade antibody recognition.J Virol. 2024 Jan 23;98(1):e0084923. doi: 10.1128/jvi.00849-23. Epub 2024 Jan 4. J Virol. 2024. PMID: 38174935 Free PMC article.
References
-
- Lemon SM, Walker C, Alter MJ, Yi M. Hepatitis C Virus. In: Knipe DM, Howley PM, editors. Fields Virology. Philadelphia, PA, USA: Lippincott Williams & Wilkins; 2007. pp. 1253–1304.
-
- Lindenbach BD, Thiel HJ, Rice CM. Flaviviridae: the viruses and their replication. In: Knipe DM, Howley PM, editors. Fields Virology. Philadelphia, PA, USA: Lippincott Williams & Wilkins; 2007. pp. 1101–1152.
-
- Drummer HE, Maerz A, Poumbourios P. Cell surface expression of functional hepatitis C virus E1 and E2 glycoproteins. FEBS Lett. 2003;546:385–390. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

