In silico partitioning and transmembrane insertion of hydrophobic peptides under equilibrium conditions
- PMID: 21861483
- PMCID: PMC3191535
- DOI: 10.1021/ja204042f
In silico partitioning and transmembrane insertion of hydrophobic peptides under equilibrium conditions
Abstract
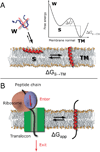
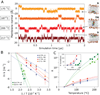
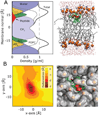
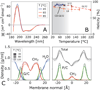
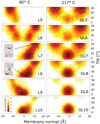
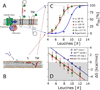
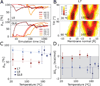
Nascent transmembrane (TM) polypeptide segments are recognized and inserted into the lipid bilayer by the cellular translocon machinery. The recognition rules, described by a biological hydrophobicity scale, correlate strongly with physical hydrophobicity scales that describe the free energy of insertion of TM helices from water. However, the exact relationship between the physical and biological scales is unknown, because solubility problems limit our ability to measure experimentally the direct partitioning of hydrophobic peptides across lipid membranes. Here we use microsecond molecular dynamics (MD) simulations in which monomeric polyleucine segments of different lengths are allowed to partition spontaneously into and out of lipid bilayers. This approach directly reveals all states populated at equilibrium. For the hydrophobic peptides studied here, only surface-bound and transmembrane-inserted helices are found. The free energy of insertion is directly obtained from the relative occupancy of these states. A water-soluble state was not observed, consistent with the general insolubility of hydrophobic peptides. The approach further allows determination of the partitioning pathways and kinetics. Surprisingly, the transfer free energy appears to be independent of temperature, which implies that surface-to-bilayer peptide insertion is a zero-entropy process. We find that the partitioning free energy of the polyleucine segments correlates strongly with values from translocon experiments but reveals a systematic shift favoring shorter peptides, suggesting that translocon-to-bilayer partitioning is not equivalent but related to spontaneous surface-to-bilayer partitioning.
Figures


References
-
- Ladokhin AS, White SH. Biochemistry. 2004;43:5782–5791. - PubMed
-
- Wimley WC, White SH. Biochemistry. 2000;39:4432–4442. - PubMed
-
- Hessa T, Kim H, Bihlmaier K, Lundin C, Boekel J, Andersson H, Nilsson I, White SH, von Heijne G. Nature. 2005;433:377–381. - PubMed
-
- Hessa T, Meindl-Beinker NM, Bernsel A, Kim H, Sato Y, Lerch-Bader M, Nilsson I, White SH, von Heijne G. Nature. 2007;450:1026–1030. - PubMed
-
- Hessa T, White SH, von Heijne G. Science. 2005;307:1427. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

