Phytoceramide and sphingoid bases derived from brewer's yeast Saccharomyces pastorianus activate peroxisome proliferator-activated receptors
- PMID: 21861924
- PMCID: PMC3176198
- DOI: 10.1186/1476-511X-10-150
Phytoceramide and sphingoid bases derived from brewer's yeast Saccharomyces pastorianus activate peroxisome proliferator-activated receptors
Abstract
Background: Peroxisome proliferator-activated receptors (PPARs) are ligand-activated transcription factors that regulate lipid and glucose metabolism. PPARα is highly expressed in the liver and controls genes involved in lipid catabolism. We previously reported that synthetic sphingolipid analogs, part of which contains shorter-length fatty acid chains than natural sphingolipids, stimulated the transcriptional activities of PPARs. Sphingosine and dihydrosphingosine (DHS) are abundant sphingoid bases, and ceramide and dihydroceramide are major ceramide species in mammals. In contrast, phytosphingosine (PHS) and DHS are the main sphingoid bases in fungi. PHS and phytoceramide exist in particular tissues such as the epidermis in mammals, and involvement of ceramide species in PPARβ activation in cultured keratinocytes has been reported. The purpose of the present study is to investigate whether natural sphingolipids with C18 fatty acid and yeast-derived sphingoid bases activate PPARs as PPAR agonists.
Method: Lipids of brewer's yeast contain PHS- and DHS-based sphingolipids. To obtain the sphingoid bases, lipids were extracted from brewer's yeast and acid-hydrolyzed. The sphingoid base fraction was purified and quantified. To assess the effects of sphingolipids on PPAR activation, luciferase reporter assay was carried out. NIH/3T3 and human hepatoma (HepG2) cells were transfected with expression vectors for PPARs and retinoid × receptors, and PPAR responsive element reporter vector. When indicated, the PPAR/Gal4 chimera system was performed to enhance the credibility of experiments. Sphingolipids were added to the cells and the dual luciferase reporter assay was performed to determine the transcriptional activity of PPARs.
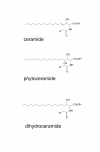
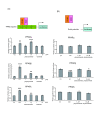
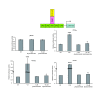
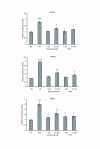
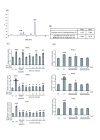
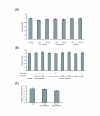
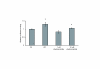
Results: We observed that phytoceramide increased the transcriptional activities of PPARs significantly, whereas ceramide and dihydroceramide did not change PPAR activities. Phytoceramide also increased transactivation of PPAR/Gal4 chimera receptors. Yeast-derived sphingoid base fraction, which contained PHS and DHS, or authentic PHS or DHS increased PPAR-dependent transcription. Additionally, phytoceramide stimulated PPARα activity in HepG2 hepatocytes, suggesting that phytoceramide activates genes regulated by PPARα.
Conclusions: Phytoceramide and yeast-derived sphingoid bases activate PPARs, whereas ceramide and dihydroceramide do not change the PPAR activity. The present findings suggest that phytoceramide acts as a PPAR ligand that would regulate PPAR-targeted genes.
Figures
Similar articles
-
Evaluation of synthetic sphingolipid analogs as ligands for peroxisome proliferator-activated receptors.Bioorg Med Chem Lett. 2009 Mar 15;19(6):1643-6. doi: 10.1016/j.bmcl.2009.02.004. Epub 2009 Feb 7. Bioorg Med Chem Lett. 2009. PMID: 19237283
-
Effects of sphingoid bases on the sphingolipidome in early keratinocyte differentiation.Exp Dermatol. 2013 Oct;22(10):677-9. doi: 10.1111/exd.12231. Exp Dermatol. 2013. PMID: 24079743
-
Do sphingoid bases interact with the peroxisome proliferator activated receptor alpha (PPAR-alpha)?Cell Signal. 2000 Jul;12(7):475-9. doi: 10.1016/s0898-6568(00)00092-9. Cell Signal. 2000. PMID: 10989283
-
International Union of Pharmacology. LXI. Peroxisome proliferator-activated receptors.Pharmacol Rev. 2006 Dec;58(4):726-41. doi: 10.1124/pr.58.4.5. Pharmacol Rev. 2006. PMID: 17132851 Review.
-
Metabolism and selected functions of sphingolipids in the yeast Saccharomyces cerevisiae.Biochim Biophys Acta. 1999 Jun 10;1438(3):305-21. doi: 10.1016/s1388-1981(99)00068-2. Biochim Biophys Acta. 1999. PMID: 10366774 Review.
Cited by
-
Egg phospholipids and cardiovascular health.Nutrients. 2015 Apr 13;7(4):2731-47. doi: 10.3390/nu7042731. Nutrients. 2015. PMID: 25871489 Free PMC article. Review.
-
New long chain bases in lipophosphonoglycan of Acanthamoeba castellanii.Lipids. 2013 Jun;48(6):639-50. doi: 10.1007/s11745-013-3794-2. Epub 2013 May 1. Lipids. 2013. PMID: 23636605
-
Structural Elucidation of Irish Ale Bioactive Polar Lipids with Antithrombotic Properties.Biomolecules. 2020 Jul 18;10(7):1075. doi: 10.3390/biom10071075. Biomolecules. 2020. PMID: 32708453 Free PMC article.
-
4,8-Sphingadienine and 4-hydroxy-8-sphingenine activate ceramide production in the skin.Lipids Health Dis. 2012 Aug 31;11:108. doi: 10.1186/1476-511X-11-108. Lipids Health Dis. 2012. PMID: 22937840 Free PMC article.
-
An integrative view of microbiome-host interactions in inflammatory bowel diseases.Cell Host Microbe. 2015 May 13;17(5):577-91. doi: 10.1016/j.chom.2015.04.008. Cell Host Microbe. 2015. PMID: 25974300 Free PMC article. Review.
References
-
- Schuler M, Ali F, Chambon C, Duteil D, Bornert JM, Tardivel A, Desvergne B, Wahli W, Chambon P, Metzger D. PGC1alpha expression is controlled in skeletal muscles by PPARbeta, whose ablation results in fiber-type switching, obesity, and type 2 diabetes. Cell Metab. 2006;4(5):407–414. doi: 10.1016/j.cmet.2006.10.003. - DOI - PubMed
-
- Man MQ, Barish GD, Schmuth M, Crumrine D, Barak Y, Chang S, Jiang Y, Evans RM, Elias PM, Feingold KR. Deficiency of PPARbeta/delta in the epidermis results in defective cutaneous permeability barrier homeostasis and increased inflammation. J Invest Dermatol. 2008;128(2):370–377. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

