5' End-independent RNase J1 endonuclease cleavage of Bacillus subtilis model RNA
- PMID: 21862575
- PMCID: PMC3186403
- DOI: 10.1074/jbc.M111.287409
5' End-independent RNase J1 endonuclease cleavage of Bacillus subtilis model RNA
Abstract
Bacillus subtilis trp leader RNA is a small (140-nucleotide) RNA that results from attenuation of trp operon transcription upon binding of the regulatory TRAP complex. Previously, endonucleolytic cleavage by ribonuclease RNase J1 in a 3'-proximal, single-stranded region was shown to be critical for initiation of trp leader RNA decay. RNase J1 is a dual-specificity enzyme, with both 5' exonucleolytic and endonucleolytic activities. Here, we provide in vivo and in vitro evidence that RNase J1 accesses its internal target site on trp leader RNA in a 5' end-independent manner. This has important implications for the role of RNase J1 in RNA decay. We also tested the involvement in trp leader RNA decay of the more recently discovered endonuclease RNase Y. Half-lives of several trp leader RNA constructs, which were designed to probe pathways of endonucleolytic versus exonucleolytic decay, were measured in an RNase Y-deficient mutant. Remarkably, the half-lives of these constructs were indistinguishable from their half-lives in an RNase J1-deficient mutant. These results suggest that lowering RNase Y concentration may affect RNA decay indirectly via an effect on RNase J1, which is thought to exist with RNase Y in a degradosome complex. To generalize our findings with trp leader RNA to other RNAs, we show that the mechanism of trp leader RNA decay is not dependent on TRAP binding.
Figures

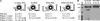



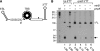
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

