GeBP/GPL transcription factors regulate a subset of CPR5-dependent processes
- PMID: 21875893
- PMCID: PMC3252139
- DOI: 10.1104/pp.111.179804
GeBP/GPL transcription factors regulate a subset of CPR5-dependent processes
Abstract
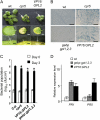
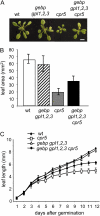
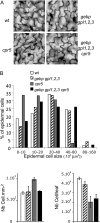
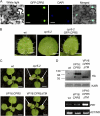
The CONSTITUTIVE EXPRESSOR OF PATHOGENESIS-RELATED GENES5 (CPR5) gene of Arabidopsis (Arabidopsis thaliana) encodes a putative membrane protein of unknown biochemical function and displays highly pleiotropic functions, particularly in pathogen responses, cell proliferation, cell expansion, and cell death. Here, we demonstrate a link between CPR5 and the GLABRA1 ENHANCER BINDING PROTEIN (GeBP) family of transcription factors. We investigated the primary role of the GeBP/GeBP-like (GPL) genes using transcriptomic analysis of the quadruple gebp gpl1,2,3 mutant and one overexpressing line that displays several cpr5-like phenotypes including dwarfism, spontaneous necrotic lesions, and increased pathogen resistance. We found that GeBP/GPLs regulate a set of genes that represents a subset of the CPR5 pathway. This subset includes genes involved in response to stress as well as cell wall metabolism. Analysis of the quintuple gebp gpl1,2,3 cpr5 mutant indicates that GeBP/GPLs are involved in the control of cell expansion in a CPR5-dependent manner but not in the control of cell proliferation. In addition, to our knowledge, we provide the first evidence that the CPR5 protein is localized in the nucleus of plant cells and that a truncated version of the protein with no transmembrane domain can trigger cpr5-like processes when fused to the VP16 constitutive transcriptional activation domain. Our results provide clues on how CPR5 and GeBP/GPLs play opposite roles in the control of cell expansion and suggest that the CPR5 protein is involved in transcription.
Figures


Similar articles
-
GeBP and GeBP-like proteins are noncanonical leucine-zipper transcription factors that regulate cytokinin response in Arabidopsis.Plant Physiol. 2008 Mar;146(3):1142-54. doi: 10.1104/pp.107.110270. Epub 2007 Dec 27. Plant Physiol. 2008. PMID: 18162594 Free PMC article.
-
Constitutive Expressor Of Pathogenesis-related Genes5 affects cell wall biogenesis and trichome development.BMC Plant Biol. 2008 May 16;8:58. doi: 10.1186/1471-2229-8-58. BMC Plant Biol. 2008. PMID: 18485217 Free PMC article.
-
GeBP, the first member of a new gene family in Arabidopsis, encodes a nuclear protein with DNA-binding activity and is regulated by KNAT1.Plant J. 2003 Jan;33(2):305-17. doi: 10.1046/j.1365-313x.2003.01622.x. Plant J. 2003. PMID: 12535344
-
Interaction of CPR5 with cell cycle regulators UVI4 and OSD1 in Arabidopsis.PLoS One. 2014 Jun 19;9(6):e100347. doi: 10.1371/journal.pone.0100347. eCollection 2014. PLoS One. 2014. PMID: 24945150 Free PMC article.
-
CPR5 is involved in cell proliferation and cell death control and encodes a novel transmembrane protein.Curr Biol. 2001 Nov 27;11(23):1891-5. doi: 10.1016/s0960-9822(01)00590-5. Curr Biol. 2001. PMID: 11728314
Cited by
-
Comparative transcriptome analysis of Zea mays upon mechanical wounding.Mol Biol Rep. 2023 Jun;50(6):5319-5343. doi: 10.1007/s11033-023-08429-x. Epub 2023 May 8. Mol Biol Rep. 2023. PMID: 37155015
-
Putative alternative translation start site-encoding nucleotides of CPR5 regulate growth and resistance.BMC Plant Biol. 2020 Jun 29;20(1):295. doi: 10.1186/s12870-020-02485-2. BMC Plant Biol. 2020. PMID: 32600419 Free PMC article.
-
Genome-Wide Identification and Expression Analysis of BrGeBP Genes Reveal Their Potential Roles in Cold and Drought Stress Tolerance in Brassica rapa.Int J Mol Sci. 2023 Sep 2;24(17):13597. doi: 10.3390/ijms241713597. Int J Mol Sci. 2023. PMID: 37686403 Free PMC article.
-
Genome-Wide Investigation and Functional Analysis Reveal That CsGeBP4 Is Required for Tea Plant Trichome Formation.Int J Mol Sci. 2023 Mar 8;24(6):5207. doi: 10.3390/ijms24065207. Int J Mol Sci. 2023. PMID: 36982281 Free PMC article.
-
STOREKEEPER RELATED1/G-Element Binding Protein (STKR1) Interacts with Protein Kinase SnRK1.Plant Physiol. 2018 Feb;176(2):1773-1792. doi: 10.1104/pp.17.01461. Epub 2017 Nov 30. Plant Physiol. 2018. PMID: 29192025 Free PMC article.
References
-
- Aguirrezabal L, Bouchier-Combaud S, Radziejwoski A, Dauzat M, Cookson SJ, Granier C. (2006) Plasticity to soil water deficit in Arabidopsis thaliana: dissection of leaf development into underlying growth dynamic and cellular variables reveals invisible phenotypes. Plant Cell Environ 29: 2216–2227 - PubMed
-
- Ahn JW, Verma R, Kim M, Lee JY, Kim YK, Bang JW, Reiter WD, Pai HS. (2006) Depletion of UDP-D-apiose/UDP-D-xylose synthases results in rhamnogalacturonan-II deficiency, cell wall thickening, and cell death in higher plants. J Biol Chem 281: 13708–13716 - PubMed
-
- Aki T, Konishi M, Kikuchi T, Fujimori T, Yoneyama T, Yanagisawa S. (2007) Distinct modulations of the hexokinase1-mediated glucose response and hexokinase1-independent processes by HYS1/CPR5 in Arabidopsis. J Exp Bot 58: 3239–3248 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

