Comparative analysis of EspF variants in inhibition of Escherichia coli phagocytosis by macrophages and inhibition of E. coli translocation through human- and bovine-derived M cells
- PMID: 21875965
- PMCID: PMC3257939
- DOI: 10.1128/IAI.00023-11
Comparative analysis of EspF variants in inhibition of Escherichia coli phagocytosis by macrophages and inhibition of E. coli translocation through human- and bovine-derived M cells
Abstract
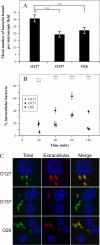
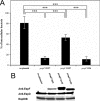
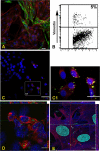
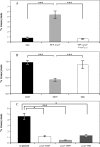
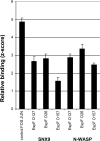
The EspF protein is secreted by the type III secretion system of enteropathogenic and enterohemorrhagic Escherichia coli (EPEC and EHEC, respectively). EspF sequences differ between EHEC O157:H7, EHEC O26:H11, and EPEC O127:H6 in terms of the number of SH3-binding polyproline-rich repeats and specific residues in these regions, as well as residues in the amino domain involved in cellular localization. EspF(O127) is important for the inhibition of phagocytosis by EPEC and also limits EPEC translocation through antigen-sampling cells (M cells). EspF(O127) has been shown to have effects on cellular organelle function and interacts with several host proteins, including N-WASP and sorting nexin 9 (SNX9). In this study, we compared the capacities of different espF alleles to inhibit (i) bacterial phagocytosis by macrophages, (ii) translocation through an M-cell coculture system, and (iii) uptake by and translocation through cultured bovine epithelial cells. The espF gene from E. coli serotype O157 (espF(O157)) allele was significantly less effective at inhibiting phagocytosis and also had reduced capacity to inhibit E. coli translocation through a human-derived in vitro M-cell coculture system in comparison to espF(O127) and espF(O26). In contrast, espF(O157) was the most effective allele at restricting bacterial uptake into and translocation through primary epithelial cells cultured from the bovine terminal rectum, the predominant colonization site of EHEC O157 in cattle and a site containing M-like cells. Although LUMIER binding assays demonstrated differences in the interactions of the EspF variants with SNX9 and N-WASP, we propose that other, as-yet-uncharacterized interactions contribute to the host-based variation in EspF activity demonstrated here.
Figures



References
-
- Bai Y., Muragaki Y., Obata K., Iwata K., Ooshima A. 1986. Immunological properties of monoclonal antibodies to human and rat prolyl 4-hydroxylase. J. Biochem. 99:1563–1570 - PubMed
-
- Barrios-Rodiles M., et al. 2005. High-throughput mapping of a dynamic signaling network in mammalian cells. Sci. STKE 307:1621–1625 - PubMed
-
- Bettelheim K. A. 2000. Role of non-O157 VTEC. Symp. Ser. Soc. Appl. Microbiol. 29:38S–50S - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

