Differentiation-dependent functional and epigenetic landscapes for cytokine genes in virus-specific CD8+ T cells
- PMID: 21876173
- PMCID: PMC3174616
- DOI: 10.1073/pnas.1112520108
Differentiation-dependent functional and epigenetic landscapes for cytokine genes in virus-specific CD8+ T cells
Abstract
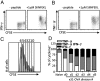
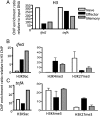
Although the simultaneous engagement of multiple effector mechanisms is thought to characterize optimal CD8(+) T-cell immunity and facilitate pathogen clearance, the differentiation pathways leading to the acquisition and maintenance of such polyfunctional activity are not well understood. Division-dependent profiles of effector molecule expression for virus-specific T cells are analyzed here by using a combination of carboxyfluorescein succinimidyl ester dilution and intracellular cytokine staining subsequent to T-cell receptor ligation. The experiments show that, although the majority of naive CD8(+) T-cell precursors are preprogrammed to produce TNF-α soon after stimulation and a proportion make both TNF-α and IL-2, the progressive acquisition of IFN-γ expression depends on continued lymphocyte proliferation. Furthermore, the extensive division characteristic of differentiation to peak effector activity is associated with the progressive dominance of IFN-γ and the concomitant loss of polyfunctional cytokine production, although this is not apparent for long-term CD8(+) T-cell memory. Such proliferation-dependent variation in cytokine production appears tied to the epigenetic signatures within the ifnG and tnfA proximal promoters. Specifically, those cytokine gene loci that are rapidly expressed following antigen stimulation at different stages of T-cell differentiation can be shown (by ChIP) to have permissive epigenetic and RNA polymerase II docking signatures. Thus, the dynamic changes in cytokine profiles for naive, effector, and memory T cells are underpinned by specific epigenetic landscapes that regulate responsiveness following T-cell receptor ligation.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



Similar articles
-
Distinct epigenetic signatures delineate transcriptional programs during virus-specific CD8(+) T cell differentiation.Immunity. 2014 Nov 20;41(5):853-65. doi: 10.1016/j.immuni.2014.11.001. Epub 2014 Nov 6. Immunity. 2014. PMID: 25517617 Free PMC article.
-
Epigenetic remodeling of the IL-2 and IFN-gamma loci in memory CD8 T cells is influenced by CD4 T cells.J Immunol. 2006 Jul 15;177(2):1062-9. doi: 10.4049/jimmunol.177.2.1062. J Immunol. 2006. PMID: 16818762
-
Effect of human bone marrow mesenchymal stromal cells on cytokine production by peripheral blood naive, memory, and effector T cells.Stem Cell Res Ther. 2015 Jan 5;6(1):3. doi: 10.1186/scrt537. Stem Cell Res Ther. 2015. PMID: 25559824 Free PMC article.
-
Epigenetic control of interferon-gamma expression in CD8 T cells.J Immunol Res. 2015;2015:849573. doi: 10.1155/2015/849573. Epub 2015 Apr 20. J Immunol Res. 2015. PMID: 25973438 Free PMC article. Review.
-
The role of epigenetics in the acquisition and maintenance of effector function in virus-specific CD8 T cells.IUBMB Life. 2010 Jul;62(7):519-26. doi: 10.1002/iub.351. IUBMB Life. 2010. PMID: 20552633 Review.
Cited by
-
T-cell memory differentiation: insights from transcriptional signatures and epigenetics.Immunology. 2013 Jul;139(3):277-84. doi: 10.1111/imm.12074. Immunology. 2013. PMID: 23347146 Free PMC article. Review.
-
Cellular and Molecular Mechanisms of CD8+ T Cell Differentiation, Dysfunction and Exhaustion.Int J Mol Sci. 2020 Oct 5;21(19):7357. doi: 10.3390/ijms21197357. Int J Mol Sci. 2020. PMID: 33027962 Free PMC article. Review.
-
Distinct epigenetic signatures delineate transcriptional programs during virus-specific CD8(+) T cell differentiation.Immunity. 2014 Nov 20;41(5):853-65. doi: 10.1016/j.immuni.2014.11.001. Epub 2014 Nov 6. Immunity. 2014. PMID: 25517617 Free PMC article.
-
Modulating DNA methylation in activated CD8+ T cells inhibits regulatory T cell-induced binding of Foxp3 to the CD8+ T Cell IL-2 promoter.J Immunol. 2015 Feb 1;194(3):990-8. doi: 10.4049/jimmunol.1401762. Epub 2014 Dec 29. J Immunol. 2015. PMID: 25548225 Free PMC article.
-
Metabolic Signaling Drives IFN-γ.Cell Metab. 2016 Nov 8;24(5):651-652. doi: 10.1016/j.cmet.2016.10.018. Cell Metab. 2016. PMID: 27829134 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

