An EMT-driven alternative splicing program occurs in human breast cancer and modulates cellular phenotype
- PMID: 21876675
- PMCID: PMC3158048
- DOI: 10.1371/journal.pgen.1002218
An EMT-driven alternative splicing program occurs in human breast cancer and modulates cellular phenotype
Abstract
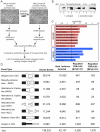
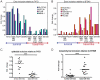
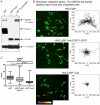
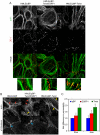
Epithelial-mesenchymal transition (EMT), a mechanism important for embryonic development, plays a critical role during malignant transformation. While much is known about transcriptional regulation of EMT, alternative splicing of several genes has also been correlated with EMT progression, but the extent of splicing changes and their contributions to the morphological conversion accompanying EMT have not been investigated comprehensively. Using an established cell culture model and RNA-Seq analyses, we determined an alternative splicing signature for EMT. Genes encoding key drivers of EMT-dependent changes in cell phenotype, such as actin cytoskeleton remodeling, regulation of cell-cell junction formation, and regulation of cell migration, were enriched among EMT-associated alternatively splicing events. Our analysis suggested that most EMT-associated alternative splicing events are regulated by one or more members of the RBFOX, MBNL, CELF, hnRNP, or ESRP classes of splicing factors. The EMT alternative splicing signature was confirmed in human breast cancer cell lines, which could be classified into basal and luminal subtypes based exclusively on their EMT-associated splicing pattern. Expression of EMT-associated alternative mRNA transcripts was also observed in primary breast cancer samples, indicating that EMT-dependent splicing changes occur commonly in human tumors. The functional significance of EMT-associated alternative splicing was tested by expression of the epithelial-specific splicing factor ESRP1 or by depletion of RBFOX2 in mesenchymal cells, both of which elicited significant changes in cell morphology and motility towards an epithelial phenotype, suggesting that splicing regulation alone can drive critical aspects of EMT-associated phenotypic changes. The molecular description obtained here may aid in the development of new diagnostic and prognostic markers for analysis of breast cancer progression.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



Comment in
-
Layers of regulation.Nat Rev Cancer. 2011 Sep 23;11(10):689. doi: 10.1038/nrc3146. Nat Rev Cancer. 2011. PMID: 21941278 No abstract available.
Similar articles
-
Splicing factor ratio as an index of epithelial-mesenchymal transition and tumor aggressiveness in breast cancer.Oncotarget. 2017 Jan 10;8(2):2423-2436. doi: 10.18632/oncotarget.13682. Oncotarget. 2017. PMID: 27911856 Free PMC article.
-
The RNA-binding protein Rbfox2: an essential regulator of EMT-driven alternative splicing and a mediator of cellular invasion.Oncogene. 2014 Feb 27;33(9):1082-92. doi: 10.1038/onc.2013.50. Epub 2013 Feb 25. Oncogene. 2014. PMID: 23435423
-
A Regulatory Axis between Epithelial Splicing Regulatory Proteins and Estrogen Receptor α Modulates the Alternative Transcriptome of Luminal Breast Cancer.Int J Mol Sci. 2022 Jul 16;23(14):7835. doi: 10.3390/ijms23147835. Int J Mol Sci. 2022. PMID: 35887187 Free PMC article.
-
Alternative splicing modulates cancer aggressiveness: role in EMT/metastasis and chemoresistance.Mol Biol Rep. 2021 Jan;48(1):897-914. doi: 10.1007/s11033-020-06094-y. Epub 2021 Jan 5. Mol Biol Rep. 2021. PMID: 33400075 Review.
-
Roles and Regulation of Epithelial Splicing Regulatory Proteins 1 and 2 in Epithelial-Mesenchymal Transition.Int Rev Cell Mol Biol. 2016;327:163-194. doi: 10.1016/bs.ircmb.2016.06.003. Epub 2016 Jul 30. Int Rev Cell Mol Biol. 2016. PMID: 27692175 Review.
Cited by
-
Altered mRNA Splicing in SMN-Depleted Motor Neuron-Like Cells.PLoS One. 2016 Oct 13;11(10):e0163954. doi: 10.1371/journal.pone.0163954. eCollection 2016. PLoS One. 2016. PMID: 27736905 Free PMC article.
-
Evolutionary dynamics of gene and isoform regulation in Mammalian tissues.Science. 2012 Dec 21;338(6114):1593-9. doi: 10.1126/science.1228186. Science. 2012. PMID: 23258891 Free PMC article.
-
The receptor AXL diversifies EGFR signaling and limits the response to EGFR-targeted inhibitors in triple-negative breast cancer cells.Sci Signal. 2013 Aug 6;6(287):ra66. doi: 10.1126/scisignal.2004155. Sci Signal. 2013. PMID: 23921085 Free PMC article.
-
Transforming Growth Factor-β-Induced RBFOX3 Inhibition Promotes Epithelial-Mesenchymal Transition of Lung Cancer Cells.Mol Cells. 2016 Aug 31;39(8):625-30. doi: 10.14348/molcells.2016.0150. Epub 2016 Jul 19. Mol Cells. 2016. PMID: 27432190 Free PMC article.
-
Adipocyte-derived exosomes may promote breast cancer progression in type 2 diabetes.Sci Signal. 2021 Nov 23;14(710):eabj2807. doi: 10.1126/scisignal.abj2807. Epub 2021 Nov 23. Sci Signal. 2021. PMID: 34813359 Free PMC article.
References
-
- Christofori G. New signals from the invasive front. Nature. 2006;441:444–450. - PubMed
-
- Yang J, Weinberg RA. Epithelial-mesenchymal transition: at the crossroads of development and tumor metastasis. Dev Cell. 2008;14:818–829. - PubMed
-
- Yilmaz M, Christofori G. EMT, the cytoskeleton, and cancer cell invasion. Cancer Metastasis Rev. 2009;28:15–33. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

