The repertoire of ICE in prokaryotes underscores the unity, diversity, and ubiquity of conjugation
- PMID: 21876676
- PMCID: PMC3158045
- DOI: 10.1371/journal.pgen.1002222
The repertoire of ICE in prokaryotes underscores the unity, diversity, and ubiquity of conjugation
Abstract
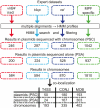
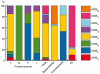
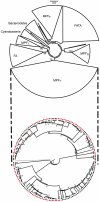
Horizontal gene transfer shapes the genomes of prokaryotes by allowing rapid acquisition of novel adaptive functions. Conjugation allows the broadest range and the highest gene transfer input per transfer event. While conjugative plasmids have been studied for decades, the number and diversity of integrative conjugative elements (ICE) in prokaryotes remained unknown. We defined a large set of protein profiles of the conjugation machinery to scan over 1,000 genomes of prokaryotes. We found 682 putative conjugative systems among all major phylogenetic clades and showed that ICEs are the most abundant conjugative elements in prokaryotes. Nearly half of the genomes contain a type IV secretion system (T4SS), with larger genomes encoding more conjugative systems. Surprisingly, almost half of the chromosomal T4SS lack co-localized relaxases and, consequently, might be devoted to protein transport instead of conjugation. This class of elements is preponderant among small genomes, is less commonly associated with integrases, and is rarer in plasmids. ICEs and conjugative plasmids in proteobacteria have different preferences for each type of T4SS, but all types exist in both chromosomes and plasmids. Mobilizable elements outnumber self-conjugative elements in both ICEs and plasmids, which suggests an extensive use of T4SS in trans. Our evolutionary analysis indicates that switch of plasmids to and from ICEs were frequent and that extant elements began to differentiate only relatively recently. According to the present results, ICEs are the most abundant conjugative elements in practically all prokaryotic clades and might be far more frequently domesticated into non-conjugative protein transport systems than previously thought. While conjugative plasmids and ICEs have different means of genomic stabilization, their mechanisms of mobility by conjugation show strikingly conserved patterns, arguing for a unitary view of conjugation in shaping the genomes of prokaryotes by horizontal gene transfer.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures




Comment in
-
A broad brush, global overview of bacterial sexuality.PLoS Genet. 2011 Aug;7(8):e1002255. doi: 10.1371/journal.pgen.1002255. Epub 2011 Aug 18. PLoS Genet. 2011. PMID: 21876683 Free PMC article. No abstract available.
References
-
- Ochman H, Lawrence JG, Groisman EA. Lateral gene transfer and the nature of bacterial innovation. Nature. 2000;405:299–304. - PubMed
-
- de la Cruz F, Davies J. Horizontal gene transfer and the origin of species: lessons from bacteria. Trends Microbiol. 2000;8:128–133. - PubMed
-
- Tettelin H, Riley D, Cattuto C, Medini D. Comparative genomics: the bacterial pan-genome. Curr Opin Microbiol. 2008;11:472–477. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

