GnRH-deficient phenotypes in humans and mice with heterozygous variants in KISS1/Kiss1
- PMID: 21880801
- PMCID: PMC3205899
- DOI: 10.1210/jc.2011-0518
GnRH-deficient phenotypes in humans and mice with heterozygous variants in KISS1/Kiss1
Abstract
Context: KISS1 is a candidate gene for GnRH deficiency.
Objective: Our objective was to identify deleterious mutations in KISS1.
Patients and methods: DNA sequencing and assessment of the effects of rare sequence variants (RSV) were conducted in 1025 probands with GnRH-deficient conditions.
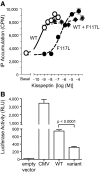
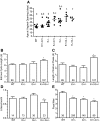
Results: Fifteen probands harbored 10 heterozygous RSV in KISS1 seen in less than 1% of control subjects. Of the variants that reside within the mature kisspeptin peptide, p.F117L (but not p.S77I, p.Q82K, p.H90D, or p.P110T) reduces inositol phosphate generation. Of the variants that lie within the coding region but outside the mature peptide, p.G35S and p.C53R (but not p.A129V) are predicted in silico to be deleterious. Of the variants that lie outside the coding region, one (g.1-3659C→T) impairs transcription in vitro, and another (c.1-7C→T) lies within the consensus Kozak sequence. Of five probands tested, four had abnormal baseline LH pulse patterns. In mice, testosterone decreases with heterozygous loss of Kiss1 and Kiss1r alleles (wild-type, 274 ± 99, to double heterozygotes, 69 ± 16 ng/dl; r(2) = 0.13; P = 0.03). Kiss1/Kiss1r double-heterozygote males have shorter anogenital distances (13.0 ± 0.2 vs. 15.6 ± 0.2 mm at P34, P < 0.001), females have longer estrous cycles (7.4 ± 0.2 vs. 5.6 ± 0.2 d, P < 0.01), and mating pairs have decreased litter frequency (0.59 ± 0.09 vs. 0.71 ± 0.06 litters/month, P < 0.04) and size (3.5 ± 0.2 vs. 5.4 ± 0.3 pups/litter, P < 0.001) compared with wild-type mice.
Conclusions: Deleterious, heterozygous RSV in KISS1 exist at a low frequency in GnRH-deficient patients as well as in the general population in presumably normal individuals. As in Kiss1(+/-)/Kiss1r(+/-) mice, heterozygous KISS1 variants in humans may work with other genetic and/or environmental factors to cause abnormal reproductive function.
Figures



Similar articles
-
[Kisspeptin and leptin in the regulation of fertility].Mol Biol (Mosk). 2015 Sep-Oct;49(5):707-15. doi: 10.7868/S0026898415050134. Mol Biol (Mosk). 2015. PMID: 26510589 Review. Russian.
-
Variants in the Kisspeptin-GnRH Pathway Modulate the Hormonal Profile and Reproductive Outcomes.DNA Cell Biol. 2020 Jun;39(6):1012-1022. doi: 10.1089/dna.2019.5165. Epub 2020 Apr 29. DNA Cell Biol. 2020. PMID: 32352843
-
Mutations of the KISS1 gene in disorders of puberty.J Clin Endocrinol Metab. 2010 May;95(5):2276-80. doi: 10.1210/jc.2009-2421. Epub 2010 Mar 17. J Clin Endocrinol Metab. 2010. PMID: 20237166 Free PMC article.
-
GnRH(1-5), a metabolite of gonadotropin-releasing hormone, enhances luteinizing hormone release via activation of kisspeptin neurons in female rats.Endocr J. 2020 Apr 28;67(4):409-418. doi: 10.1507/endocrj.EJ19-0444. Epub 2020 Jan 15. Endocr J. 2020. PMID: 31941848
-
The kisspeptin signaling pathway and its role in human isolated GnRH deficiency.Mol Cell Endocrinol. 2011 Oct 22;346(1-2):29-36. doi: 10.1016/j.mce.2011.05.043. Epub 2011 Jun 17. Mol Cell Endocrinol. 2011. PMID: 21704672 Free PMC article. Review.
Cited by
-
Expert consensus document: European Consensus Statement on congenital hypogonadotropic hypogonadism--pathogenesis, diagnosis and treatment.Nat Rev Endocrinol. 2015 Sep;11(9):547-64. doi: 10.1038/nrendo.2015.112. Epub 2015 Jul 21. Nat Rev Endocrinol. 2015. PMID: 26194704 Review.
-
Mutations in FGF17, IL17RD, DUSP6, SPRY4, and FLRT3 are identified in individuals with congenital hypogonadotropic hypogonadism.Am J Hum Genet. 2013 May 2;92(5):725-43. doi: 10.1016/j.ajhg.2013.04.008. Am J Hum Genet. 2013. PMID: 23643382 Free PMC article.
-
GPR54-dependent stimulation of luteinizing hormone secretion by neurokinin B in prepubertal rats.PLoS One. 2012;7(9):e44344. doi: 10.1371/journal.pone.0044344. Epub 2012 Sep 28. PLoS One. 2012. PMID: 23028524 Free PMC article.
-
Kisspeptin deficiency leads to abnormal adrenal glands and excess steroid hormone secretion.Hum Mol Genet. 2020 Dec 18;29(20):3443-3450. doi: 10.1093/hmg/ddaa215. Hum Mol Genet. 2020. PMID: 33089319 Free PMC article.
-
Congenital hypogonadotropic hypogonadism and constitutional delay of growth and puberty have distinct genetic architectures.Eur J Endocrinol. 2018 Apr;178(4):377-388. doi: 10.1530/EJE-17-0568. Epub 2018 Feb 1. Eur J Endocrinol. 2018. PMID: 29419413 Free PMC article.
References
-
- Dodé C, Levilliers J, Dupont JM, De Paepe A, Le Dû N, Soussi-Yanicostas N, Coimbra RS, Delmaghani S, Compain-Nouaille S, Baverel F, Pêcheux C, Le Tessier D, Cruaud C, Delpech M, Speleman F, Vermeulen S, Amalfitano A, Bachelot Y, Bouchard P, Cabrol S, Carel JC, Delemarre-van de Waal H, Goulet-Salmon B, Kottler ML, Richard O, Sanchez-Franco F, Saura R, Young J, Petit C, Hardelin JP. 2003. Loss-of-function mutations in FGFR1 cause autosomal dominant Kallmann syndrome. Nat Genet 33:463–465 - PubMed
-
- Falardeau J, Chung WC, Beenken A, Raivio T, Plummer L, Sidis Y, Jacobson-Dickman EE, Eliseenkova AV, Ma J, Dwyer A, Quinton R, Na S, Hall JE, Huot C, Alois N, Pearce SH, Cole LW, Hughes V, Mohammadi M, Tsai P, Pitteloud N. 2008. Decreased FGF8 signaling causes deficiency of gonadotropin-releasing hormone in humans and mice. J Clin Invest 118:2822–2831 - PMC - PubMed
-
- Dodé C, Teixeira L, Levilliers J, Fouveaut C, Bouchard P, Kottler ML, Lespinasse J, Lienhardt-Roussie A, Mathieu M, Moerman A, Morgan G, Murat A, Toublanc JE, Wolczynski S, Delpech M, Petit C, Young J, Hardelin JP. 2006. Kallmann syndrome: mutations in the genes encoding prokineticin-2 and prokineticin receptor-2. PLoS Genet 2:e175. - PMC - PubMed
-
- Topaloglu AK, Reimann F, Guclu M, Yalin AS, Kotan LD, Porter KM, Serin A, Mungan NO, Cook JR, Ozbek MN, Imamoglu S, Akalin NS, Yuksel B, O'Rahilly S, Semple RK. 2009. TAC3 and TACR3 mutations in familial hypogonadotropic hypogonadism reveal a key role for Neurokinin B in the central control of reproduction. Nat Genet 41:354–358 - PMC - PubMed
-
- de Roux N, Young J, Misrahi M, Genet R, Chanson P, Schaison G, Milgrom E. 1997. A family with hypogonadotropic hypogonadism and mutations in the gonadotropin-releasing hormone receptor. N Engl J Med 337:1597–1602 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

