Molecular dissection of the roles of phytochrome in photoperiodic flowering in rice
- PMID: 21880933
- PMCID: PMC3252176
- DOI: 10.1104/pp.111.181792
Molecular dissection of the roles of phytochrome in photoperiodic flowering in rice
Abstract
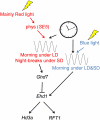
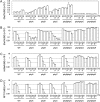
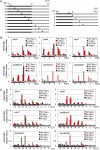
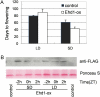
Phytochromes mediate the photoperiodic control of flowering in rice (Oryza sativa), a short-day plant. Recent molecular genetics studies have revealed a genetic network that enables the critical daylength response of florigen gene expression. Analyses using a rice phytochrome chromophore-deficient mutant, photoperiod sensitivity5, have so far revealed that within this network, phytochromes are required for expression of Grain number, plant height and heading date7 (Ghd7), a floral repressor gene in rice. There are three phytochrome genes in rice, but the roles of each phytochrome family member in daylength response have not previously been defined. Here, we revealed multiple action points for each phytochrome in the critical daylength response of florigen expression by using single and double phytochrome mutant lines of rice. Our results show that either phyA alone or a genetic combination of phyB and phyC can induce Ghd7 mRNA, whereas phyB alone causes some reduction in levels of Ghd7 mRNA. Moreover, phyB and phyA can affect Ghd7 activity and Early heading date1 (a floral inducer) activity in the network, respectively. Therefore, each phytochrome gene of rice has distinct roles, and all of the phytochrome actions coordinately control the critical daylength response of florigen expression in rice.
Figures

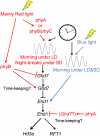
References
-
- Borthwick HA. (1964) Phytochrome action and its time displays. Am Nat 98: 347–355
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

