Divalent metal ion differentially regulates the sequential nicking reactions of the GIY-YIG homing endonuclease I-BmoI
- PMID: 21887323
- PMCID: PMC3161791
- DOI: 10.1371/journal.pone.0023804
Divalent metal ion differentially regulates the sequential nicking reactions of the GIY-YIG homing endonuclease I-BmoI
Abstract
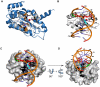
Homing endonucleases are site-specific DNA endonucleases that function as mobile genetic elements by introducing double-strand breaks or nicks at defined locations. Of the major families of homing endonucleases, the modular GIY-YIG endonucleases are least understood in terms of mechanism. The GIY-YIG homing endonuclease I-BmoI generates a double-strand break by sequential nicking reactions during which the single active site of the GIY-YIG nuclease domain must undergo a substantial reorganization. Here, we show that divalent metal ion plays a significant role in regulating the two independent nicking reactions by I-BmoI. Rate constant determination for each nicking reaction revealed that limiting divalent metal ion has a greater impact on the second strand than the first strand nicking reaction. We also show that substrate mutations within the I-BmoI cleavage site can modulate the first strand nicking reaction over a 314-fold range. Additionally, in-gel DNA footprinting with mutant substrates and modeling of an I-BmoI-substrate complex suggest that amino acid contacts to a critical GC-2 base pair are required to induce a bottom-strand distortion that likely directs conformational changes for reaction progress. Collectively, our data implies mechanistic roles for divalent metal ion and substrate bases, suggesting that divalent metal ion facilitates the re-positioning of the GIY-YIG nuclease domain between sequential nicking reactions.
Conflict of interest statement
Figures





Similar articles
-
The monomeric GIY-YIG homing endonuclease I-BmoI uses a molecular anchor and a flexible tether to sequentially nick DNA.Nucleic Acids Res. 2013 May 1;41(10):5413-27. doi: 10.1093/nar/gkt186. Epub 2013 Apr 4. Nucleic Acids Res. 2013. PMID: 23558745 Free PMC article.
-
Strand-specific contacts and divalent metal ion regulate double-strand break formation by the GIY-YIG homing endonuclease I-BmoI.J Mol Biol. 2007 Nov 23;374(2):306-21. doi: 10.1016/j.jmb.2007.09.027. Epub 2007 Sep 16. J Mol Biol. 2007. PMID: 17936302
-
Distance determination by GIY-YIG intron endonucleases: discrimination between repression and cleavage functions.Nucleic Acids Res. 2006 Mar 31;34(6):1755-64. doi: 10.1093/nar/gkl079. Print 2006. Nucleic Acids Res. 2006. PMID: 16582101 Free PMC article.
-
Homing endonuclease structure and function.Q Rev Biophys. 2005 Feb;38(1):49-95. doi: 10.1017/S0033583505004063. Epub 2005 Dec 9. Q Rev Biophys. 2005. PMID: 16336743 Review.
-
Catalytic mechanisms of restriction and homing endonucleases.Biochemistry. 2002 Nov 26;41(47):13851-60. doi: 10.1021/bi020467h. Biochemistry. 2002. PMID: 12437341 Review.
Cited by
-
The monomeric GIY-YIG homing endonuclease I-BmoI uses a molecular anchor and a flexible tether to sequentially nick DNA.Nucleic Acids Res. 2013 May 1;41(10):5413-27. doi: 10.1093/nar/gkt186. Epub 2013 Apr 4. Nucleic Acids Res. 2013. PMID: 23558745 Free PMC article.
-
Monomeric site-specific nucleases for genome editing.Proc Natl Acad Sci U S A. 2012 May 22;109(21):8061-6. doi: 10.1073/pnas.1117984109. Epub 2012 May 7. Proc Natl Acad Sci U S A. 2012. PMID: 22566637 Free PMC article.
-
Plant Organellar MSH1 Is a Displacement Loop-Specific Endonuclease.Plant Cell Physiol. 2024 May 14;65(4):560-575. doi: 10.1093/pcp/pcad112. Plant Cell Physiol. 2024. PMID: 37756637 Free PMC article.
-
Spectral analysis of naturally occurring methylxanthines (theophylline, theobromine and caffeine) binding with DNA.PLoS One. 2012;7(12):e50019. doi: 10.1371/journal.pone.0050019. Epub 2012 Dec 7. PLoS One. 2012. PMID: 23236361 Free PMC article.
-
RecA-dependent programmable endonuclease Ref cleaves DNA in two distinct steps.Nucleic Acids Res. 2014 Apr;42(6):3871-83. doi: 10.1093/nar/gkt1342. Epub 2013 Dec 26. Nucleic Acids Res. 2014. PMID: 24371286 Free PMC article.
References
-
- Belfort M, Derbyshire V, Cousineau B, Lambowitz A. Mobile introns: pathways and proteins. In: Craig N, Craigie R, Gellert M, Lambowitz A, editors. Mobile DNA II. New York: ASM Press; 2002. pp. 761–783.
-
- Dujon B, Belfort M, Butow RA, Jacq C, Lemieux C, et al. Mobile introns: definition of terms and recommended nomenclature. Gene. 1989;82:115–118. - PubMed
-
- Bell-Pedersen D, Quirk SM, Aubrey M, Belfort M. A site-specific endonuclease and co-conversion of flanking exons associated with the mobile td intron of phage T4. Gene. 1989;82:119–126. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

