Evolution and functional diversification of fructose bisphosphate aldolase genes in photosynthetic marine diatoms
- PMID: 21903677
- PMCID: PMC3245544
- DOI: 10.1093/molbev/msr223
Evolution and functional diversification of fructose bisphosphate aldolase genes in photosynthetic marine diatoms
Abstract
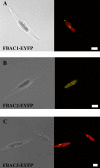
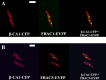
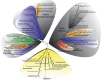
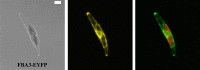
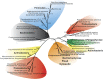
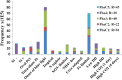
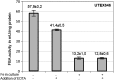
Diatoms and other chlorophyll-c containing, or chromalveolate, algae are among the most productive and diverse phytoplankton in the ocean. Evolutionarily, chlorophyll-c algae are linked through common, although not necessarily monophyletic, acquisition of plastid endosymbionts of red as well as most likely green algal origin. There is also strong evidence for a relatively high level of lineage-specific bacterial gene acquisition within chromalveolates. Therefore, analyses of gene content and derivation in chromalveolate taxa have indicated particularly diverse origins of their overall gene repertoire. As a single group of functionally related enzymes spanning two distinct gene families, fructose 1,6-bisphosphate aldolases (FBAs) illustrate the influence on core biochemical pathways of specific evolutionary associations among diatoms and other chromalveolates with various plastid-bearing and bacterial endosymbionts. Protein localization and activity, gene expression, and phylogenetic analyses indicate that the pennate diatom Phaeodactylum tricornutum contains five FBA genes with very little overall functional overlap. Three P. tricornutum FBAs, one class I and two class II, are plastid localized, and each appears to have a distinct evolutionary origin as well as function. Class I plastid FBA appears to have been acquired by chromalveolates from a red algal endosymbiont, whereas one copy of class II plastid FBA is likely to have originated from an ancient green algal endosymbiont. The other copy appears to be the result of a chromalveolate-specific gene duplication. Plastid FBA I and chromalveolate-specific class II plastid FBA are localized in the pyrenoid region of the chloroplast where they are associated with β-carbonic anhydrase, which is known to play a significant role in regulation of the diatom carbon concentrating mechanism. The two pyrenoid-associated FBAs are distinguished by contrasting gene expression profiles under nutrient limiting compared with optimal CO2 fixation conditions, suggestive of a distinct specialized function for each. Cytosolically localized FBAs in P. tricornutum likely play a role in glycolysis and cytoskeleton function and seem to have originated from the stramenopile host cell and from diatom-specific bacterial gene transfer, respectively.
Figures



Similar articles
-
Gene replacement of fructose-1,6-bisphosphate aldolase supports the hypothesis of a single photosynthetic ancestor of chromalveolates.Eukaryot Cell. 2004 Oct;3(5):1169-75. doi: 10.1128/EC.3.5.1169-1175.2004. Eukaryot Cell. 2004. PMID: 15470245 Free PMC article.
-
Thylakoid luminal θ-carbonic anhydrase critical for growth and photosynthesis in the marine diatom Phaeodactylum tricornutum.Proc Natl Acad Sci U S A. 2016 Aug 30;113(35):9828-33. doi: 10.1073/pnas.1603112113. Epub 2016 Aug 16. Proc Natl Acad Sci U S A. 2016. PMID: 27531955 Free PMC article.
-
The peculiar distribution of class I and class II aldolases in diatoms and in red algae.Curr Genet. 2005 Dec;48(6):389-400. doi: 10.1007/s00294-005-0033-2. Epub 2005 Nov 5. Curr Genet. 2005. PMID: 16273368
-
Molecular aspects of the biophysical CO2-concentrating mechanism and its regulation in marine diatoms.J Exp Bot. 2017 Jun 1;68(14):3763-3772. doi: 10.1093/jxb/erx173. J Exp Bot. 2017. PMID: 28633304 Review.
-
Mechanisms of carbon dioxide acquisition and CO2 sensing in marine diatoms: a gateway to carbon metabolism.Philos Trans R Soc Lond B Biol Sci. 2017 Sep 5;372(1728):20160403. doi: 10.1098/rstb.2016.0403. Philos Trans R Soc Lond B Biol Sci. 2017. PMID: 28717013 Free PMC article. Review.
Cited by
-
Genome sequence and comparative analysis of a putative entomopathogenic Serratia isolated from Caenorhabditis briggsae.BMC Genomics. 2015 Jul 18;16(1):531. doi: 10.1186/s12864-015-1697-8. BMC Genomics. 2015. PMID: 26187596 Free PMC article.
-
Short-term response to iron resupply in an iron-limited open ocean diatom reveals rapid decay of iron-responsive transcripts.PLoS One. 2023 Jan 24;18(1):e0280827. doi: 10.1371/journal.pone.0280827. eCollection 2023. PLoS One. 2023. PMID: 36693065 Free PMC article.
-
Reevaluating the green contribution to diatom genomes.Genome Biol Evol. 2012;4(7):683-8. doi: 10.1093/gbe/evs053. Epub 2012 Jun 7. Genome Biol Evol. 2012. PMID: 22684208 Free PMC article.
-
Experimental evidence and isotopomer analysis of mixotrophic glucose metabolism in the marine diatom Phaeodactylum tricornutum.Microb Cell Fact. 2013 Nov 14;12:109. doi: 10.1186/1475-2859-12-109. Microb Cell Fact. 2013. PMID: 24228629 Free PMC article.
-
The intracellular distribution of inorganic carbon fixing enzymes does not support the presence of a C4 pathway in the diatom Phaeodactylum tricornutum.Photosynth Res. 2018 Aug;137(2):263-280. doi: 10.1007/s11120-018-0500-5. Epub 2018 Mar 23. Photosynth Res. 2018. PMID: 29572588
References
-
- Allen AE, Dupont CL, Obornik M, et al. (11 co-authors) Evolution and metabolic significance of the urea cycle in photosynthetic diatoms. Nature. 2011;473:203–207. - PubMed
-
- Allen AE, Vardi A, Bowler C. An ecological and evolutionary context for integrated nitrogen metabolism and related signaling pathways in marine diatoms. Cur Opin Plant Biol. 2006;9:264–273. - PubMed
-
- Anisimova M, Gascuel O. Approximate likelihood-ratio test for branches: a fast, accurate, and powerful alternative. Syst Biol. 2006;55:539–552. - PubMed
-
- Anita N. Comparative studies on aldolase activity in marine planktonic algae and their evolutionary significance. J Phycol. 1967;3:81–85. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

