Roles of mutation and selection in speciation: from Hugo de Vries to the modern genomic era
- PMID: 21903731
- PMCID: PMC3227404
- DOI: 10.1093/gbe/evr028
Roles of mutation and selection in speciation: from Hugo de Vries to the modern genomic era
Abstract
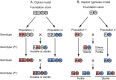
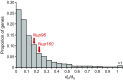
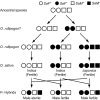
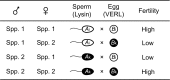
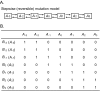
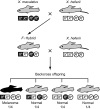
One of the most important problems in evolutionary biology is to understand how new species are generated in nature. In the past, it was difficult to study this problem because our lifetime is too short to observe the entire process of speciation. In recent years, however, molecular and genomic techniques have been developed for identifying and studying the genes involved in speciation. Using these techniques, many investigators have already obtained new findings. At present, however, the results obtained are complex and quite confusing. We have therefore attempted to understand these findings coherently with a historical perspective and clarify the roles of mutation and natural selection in speciation. We have first indicated that the root of the currently burgeoning field of plant genomics goes back to Hugo de Vries, who proposed the mutation theory of evolution more than a century ago and that he unknowingly found the importance of polyploidy and chromosomal rearrangements in plant speciation. We have then shown that the currently popular Dobzhansky-Muller model of evolution of reproductive isolation is only one of many possible mechanisms. Some of them are Oka's model of duplicate gene mutations, multiallelic speciation, mutation-rescue model, segregation-distorter gene model, heterochromatin-associated speciation, single-locus model, etc. The occurrence of speciation also depends on the reproductive system, population size, bottleneck effects, and environmental factors, such as temperature and day length. Some authors emphasized the importance of natural selection to speed up speciation, but mutation is crucial in speciation because reproductive barriers cannot be generated without mutations.
Figures
Similar articles
-
Viviparity-driven conflict: more to speciation than meets the fly.Ann N Y Acad Sci. 2008;1133:126-48. doi: 10.1196/annals.1438.006. Ann N Y Acad Sci. 2008. PMID: 18559818 Review.
-
Review. The strength and genetic basis of reproductive isolating barriers in flowering plants.Philos Trans R Soc Lond B Biol Sci. 2008 Sep 27;363(1506):3009-21. doi: 10.1098/rstb.2008.0064. Philos Trans R Soc Lond B Biol Sci. 2008. PMID: 18579478 Free PMC article. Review.
-
Muller "Elements" in Drosophila: How the Search for the Genetic Basis for Speciation Led to the Birth of Comparative Genomics.Genetics. 2018 Sep;210(1):3-13. doi: 10.1534/genetics.118.301084. Genetics. 2018. PMID: 30166445 Free PMC article.
-
Evolution, selection and isolation: a genomic view of speciation in fungal plant pathogens.New Phytol. 2013 Sep;199(4):895-907. doi: 10.1111/nph.12374. Epub 2013 Jun 19. New Phytol. 2013. PMID: 23782262 Review.
-
Interpreting the genomic landscape of speciation: a road map for finding barriers to gene flow.J Evol Biol. 2017 Aug;30(8):1450-1477. doi: 10.1111/jeb.13047. J Evol Biol. 2017. PMID: 28786193 Review.
Cited by
-
Global Evolutionary Analysis of 11 Gene Families Part of Reactive Oxygen Species (ROS) Gene Network in Four Eucalyptus Species.Antioxidants (Basel). 2020 Mar 21;9(3):257. doi: 10.3390/antiox9030257. Antioxidants (Basel). 2020. PMID: 32245199 Free PMC article.
-
Deep sequencing of wheat sRNA transcriptome reveals distinct temporal expression pattern of miRNAs in response to heat, light and UV.Sci Rep. 2016 Dec 22;6:39373. doi: 10.1038/srep39373. Sci Rep. 2016. PMID: 28004741 Free PMC article.
-
Environmental Epigenetics and a Unified Theory of the Molecular Aspects of Evolution: A Neo-Lamarckian Concept that Facilitates Neo-Darwinian Evolution.Genome Biol Evol. 2015 Apr 26;7(5):1296-302. doi: 10.1093/gbe/evv073. Genome Biol Evol. 2015. PMID: 25917417 Free PMC article. Review.
-
Pollen-mediated gene flow from glyphosate-resistant common waterhemp (Amaranthus rudis Sauer): consequences for the dispersal of resistance genes.Sci Rep. 2017 Mar 22;7:44913. doi: 10.1038/srep44913. Sci Rep. 2017. PMID: 28327669 Free PMC article.
-
Coupled evolutionary rates shape a Hawaiian insect-symbiont system.BMC Genomics. 2025 Apr 3;26(1):336. doi: 10.1186/s12864-025-11514-z. BMC Genomics. 2025. PMID: 40181281 Free PMC article.
References
-
- Adams KL, Wendel JF. Polyploidy and genome evolution in plants. Curr Opin Plant Biol. 2005;8:135–141. - PubMed
-
- Allen GE. Hugo de Vries and the reception of the “mutation theory”. J Hist Biol. 1969;2:55–87.
-
- Badaeva ED, et al. Chromosomal rearrangements in wheat: their types and distribution. Genome. 2007;50:907–926. - PubMed
-
- Bateson W. Heredity and variation in modern lights. In: Seward AC, editor. Darwin and modern science. Cambridge: Cambridge University Press; 1909. pp. 85–101.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

