Structure of fumarate hydratase from Rickettsia prowazekii, the agent of typhus and suspected relative of the mitochondria
- PMID: 21904061
- PMCID: PMC3169413
- DOI: 10.1107/S174430911102690X
Structure of fumarate hydratase from Rickettsia prowazekii, the agent of typhus and suspected relative of the mitochondria
Abstract
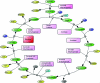
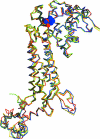
Rickettsiae are obligate intracellular parasites of eukaryotic cells that are the causative agents responsible for spotted fever and typhus. Their small genome (about 800 protein-coding genes) is highly conserved across species and has been postulated as the ancestor of the mitochondria. No genes that are required for glycolysis are found in the Rickettsia prowazekii or mitochondrial genomes, but a complete set of genes encoding components of the tricarboxylic acid cycle and the respiratory-chain complex is found in both. A 2.4 Å resolution crystal structure of R. prowazekii fumarate hydratase, an enzyme catalyzing the third step of the tricarboxylic acid cycle pathway that ultimately converts phosphoenolpyruvate into succinyl-CoA, has been solved. A structure alignment with human mitochondrial fumarate hydratase highlights the close similarity between R. prowazekii and mitochondrial enzymes.
Figures

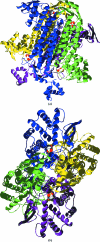

References
-
- Andersson, S. G., Zomorodipour, A., Andersson, J. O., Sicheritz-Pontén, T., Alsmark, U. C., Podowski, R. M., Näslund, A. K., Eriksson, A. S., Winkler, H. H. & Kurland, C. G. (1998). Nature (London), 396, 133–140. - PubMed
-
- Azad, A. F. (2007). Clin. Infect. Dis. 45, S52–S55.
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources

