Identification of IL6R and chromosome 11q13.5 as risk loci for asthma
- PMID: 21907864
- PMCID: PMC3517659
- DOI: 10.1016/S0140-6736(11)60874-X
Identification of IL6R and chromosome 11q13.5 as risk loci for asthma
Abstract
Background: We aimed to identify novel genetic variants affecting asthma risk, since these might provide novel insights into molecular mechanisms underlying the disease.
Methods: We did a genome-wide association study (GWAS) in 2669 physician-diagnosed asthmatics and 4528 controls from Australia. Seven loci were prioritised for replication after combining our results with those from the GABRIEL consortium (n=26,475), and these were tested in an additional 25,358 independent samples from four in-silico cohorts. Quantitative multi-marker scores of genetic load were constructed on the basis of results from the GABRIEL study and tested for association with asthma in our Australian GWAS dataset.
Findings: Two loci were confirmed to associate with asthma risk in the replication cohorts and reached genome-wide significance in the combined analysis of all available studies (n=57,800): rs4129267 (OR 1·09, combined p=2·4×10(-8)) in the interleukin-6 receptor (IL6R) gene and rs7130588 (OR 1·09, p=1·8×10(-8)) on chromosome 11q13.5 near the leucine-rich repeat containing 32 gene (LRRC32, also known as GARP). The 11q13.5 locus was significantly associated with atopic status among asthmatics (OR 1·33, p=7×10(-4)), suggesting that it is a risk factor for allergic but not non-allergic asthma. Multi-marker association results are consistent with a highly polygenic contribution to asthma risk, including loci with weak effects that might be shared with other immune-related diseases, such as NDFIP1, HLA-B, LPP, and BACH2.
Interpretation: The IL6R association further supports the hypothesis that cytokine signalling dysregulation affects asthma risk, and raises the possibility that an IL6R antagonist (tocilizumab) may be effective to treat the disease, perhaps in a genotype-dependent manner. Results for the 11q13.5 locus suggest that it directly increases the risk of allergic sensitisation which, in turn, increases the risk of subsequent development of asthma. Larger or more functionally focused studies are needed to characterise the many loci with modest effects that remain to be identified for asthma.
Funding: National Health and Medical Research Council of Australia. A full list of funding sources is provided in the webappendix.
Copyright © 2011 Elsevier Ltd. All rights reserved.
Figures
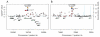
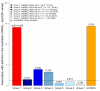
Comment in
-
Successfully mapping novel asthma loci by GWAS.Lancet. 2011 Sep 10;378(9795):967-8. doi: 10.1016/S0140-6736(11)61422-0. Lancet. 2011. PMID: 21907849 No abstract available.
References
-
- Moffatt MF, Kabesch M, Liang L, Dixon AL, Strachan D, Heath S, et al. Genetic variants regulating ORMDL3 expression contribute to the risk of childhood asthma. Nature. 2007 Jul 26;448(7152):470–3. - PubMed
-
- Sleiman PM, Flory J, Imielinski M, Bradfield JP, Annaiah K, Willis-Owen SA, et al. Variants of DENND1B associated with asthma in children. N Engl J Med. 2010 Jan 7;362(1):36–44. - PubMed
-
- Gudbjartsson DF, Bjornsdottir US, Halapi E, Helgadottir A, Sulem P, Jonsdottir GM, et al. Sequence variants affecting eosinophil numbers associate with asthma and myocardial infarction. Nat Genet. 2009 Mar;41(3):342–7. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- R01 AA007535/AA/NIAAA NIH HHS/United States
- DA12854/DA/NIDA NIH HHS/United States
- DH_/Department of Health/United Kingdom
- AA17688/AA/NIAAA NIH HHS/United States
- R01 AA013321/AA/NIAAA NIH HHS/United States
- AA07728/AA/NIAAA NIH HHS/United States
- R01 DA012854/DA/NIDA NIH HHS/United States
- R01 AA010249/AA/NIAAA NIH HHS/United States
- G0500539/MRC_/Medical Research Council/United Kingdom
- WT_/Wellcome Trust/United Kingdom
- AA14041/AA/NIAAA NIH HHS/United States
- AA13326/AA/NIAAA NIH HHS/United States
- R01 AA013320/AA/NIAAA NIH HHS/United States
- N02-HL-6-4278/HL/NHLBI NIH HHS/United States
- N01-HC-25195/HC/NHLBI NIH HHS/United States
- R01 AA014041/AA/NIAAA NIH HHS/United States
- K05 AA017688/AA/NIAAA NIH HHS/United States
- R01 MH066206/MH/NIMH NIH HHS/United States
- MOP-82893/CAPMC/ CIHR/Canada
- AA13320/AA/NIAAA NIH HHS/United States
- R01 HL087679/HL/NHLBI NIH HHS/United States
- P60 AA011998/AA/NIAAA NIH HHS/United States
- N02 HL064278/HL/NHLBI NIH HHS/United States
- P50 AA011998/AA/NIAAA NIH HHS/United States
- AA13321/AA/NIAAA NIH HHS/United States
- AA10248/AA/NIAAA NIH HHS/United States
- R01 HD042157/HD/NICHD NIH HHS/United States
- N01 HC025195/HC/NHLBI NIH HHS/United States
- R01 AA013326/AA/NIAAA NIH HHS/United States
- R56 DA012854/DA/NIDA NIH HHS/United States
- 1RL1MH083268-01/MH/NIMH NIH HHS/United States
- P30 NS062691/NS/NINDS NIH HHS/United States
- MH66206/MH/NIMH NIH HHS/United States
- AA11998/AA/NIAAA NIH HHS/United States
- R01D0042157-01A/PHS HHS/United States
- 5R01HL087679-02/HL/NHLBI NIH HHS/United States
- R37 AA007728/AA/NIAAA NIH HHS/United States
- RL1 MH083268/MH/NIMH NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

