Molecular basis of α1-antitrypsin deficiency revealed by the structure of a domain-swapped trimer
- PMID: 21909074
- PMCID: PMC3185345
- DOI: 10.1038/embor.2011.171
Molecular basis of α1-antitrypsin deficiency revealed by the structure of a domain-swapped trimer
Abstract
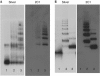
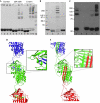
α(1)-Antitrypsin (α1AT) deficiency is a disease with multiple manifestations, including cirrhosis and emphysema, caused by the accumulation of stable polymers of mutant protein in the endoplasmic reticulum of hepatocytes. However, the molecular basis of misfolding and polymerization remain unknown. We produced and crystallized a trimeric form of α1AT that is recognized by an antibody specific for the pathological polymer. Unexpectedly, this structure reveals a polymeric linkage mediated by domain swapping the carboxy-terminal 34 residues. Disulphide-trapping and antibody-binding studies further demonstrate that runaway C-terminal domain swapping, rather than the s4A/s5A domain swap previously proposed, underlies polymerization of the common Z-mutant of α1AT in vivo.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures

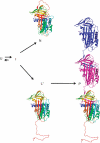
Comment in
-
The structural diversity in α1-antitrypsin misfolding.EMBO Rep. 2011 Sep 30;12(10):983-4. doi: 10.1038/embor.2011.187. EMBO Rep. 2011. PMID: 21921939 Free PMC article. No abstract available.
References
-
- Davies MJ, Lomas DA (2008) The molecular aetiology of the serpinopathies. Int J Biochem Cell Biol 40: 1273–1286 - PubMed
-
- DeLano WL (2002) The PyMOL Molecular Graphics System (on World Wide Web) http://www.pymol.org
-
- Emsley P, Cowtan K (2004) Coot: model-building tools for molecular graphics. Acta Crystallogr D Biol Crystallogr 60: 2126–2132 - PubMed
-
- Gooptu B, Lomas DA (2009) Conformational pathology of the serpins: themes, variations, and therapeutic strategies. Annu Rev Biochem 78: 147–176 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

