Thermodynamically stable RNA three-way junction for constructing multifunctional nanoparticles for delivery of therapeutics
- PMID: 21909084
- PMCID: PMC3189281
- DOI: 10.1038/nnano.2011.105
Thermodynamically stable RNA three-way junction for constructing multifunctional nanoparticles for delivery of therapeutics
Abstract
RNA nanoparticles have applications in the treatment of cancers and viral infection; however, the instability of RNA nanoparticles has hindered their development for therapeutic applications. The lack of covalent linkage or crosslinking in nanoparticles causes dissociation in vivo. Here we show that the packaging RNA of bacteriophage phi29 DNA packaging motor can be assembled from 3-6 pieces of RNA oligomers without the use of metal salts. Each RNA oligomer contains a functional module that can be a receptor-binding ligand, aptamer, short interfering RNA or ribozyme. When mixed together, they self-assemble into thermodynamically stable tri-star nanoparticles with a three-way junction core. These nanoparticles are resistant to 8 M urea denaturation, are stable in serum and remain intact at extremely low concentrations. The modules remain functional in vitro and in vivo, suggesting that the three-way junction core can be used as a platform for building a variety of multifunctional nanoparticles. We studied 25 different three-way junction motifs in biological RNA and found only one other motif that shares characteristics similar to the three-way junction of phi29 pRNA.
Figures





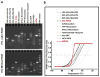
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

