Identification of single- and multiple-class specific signature genes from gene expression profiles by group marker index
- PMID: 21909426
- PMCID: PMC3164723
- DOI: 10.1371/journal.pone.0024259
Identification of single- and multiple-class specific signature genes from gene expression profiles by group marker index
Abstract
Informative genes from microarray data can be used to construct prediction model and investigate biological mechanisms. Differentially expressed genes, the main targets of most gene selection methods, can be classified as single- and multiple-class specific signature genes. Here, we present a novel gene selection algorithm based on a Group Marker Index (GMI), which is intuitive, of low-computational complexity, and efficient in identification of both types of genes. Most gene selection methods identify only single-class specific signature genes and cannot identify multiple-class specific signature genes easily. Our algorithm can detect de novo certain conditions of multiple-class specificity of a gene and makes use of a novel non-parametric indicator to assess the discrimination ability between classes. Our method is effective even when the sample size is small as well as when the class sizes are significantly different. To compare the effectiveness and robustness we formulate an intuitive template-based method and use four well-known datasets. We demonstrate that our algorithm outperforms the template-based method in difficult cases with unbalanced distribution. Moreover, the multiple-class specific genes are good biomarkers and play important roles in biological pathways. Our literature survey supports that the proposed method identifies unique multiple-class specific marker genes (not reported earlier to be related to cancer) in the Central Nervous System data. It also discovers unique biomarkers indicating the intrinsic difference between subtypes of lung cancer. We also associate the pathway information with the multiple-class specific signature genes and cross-reference to published studies. We find that the identified genes participate in the pathways directly involved in cancer development in leukemia data. Our method gives a promising way to find genes that can involve in pathways of multiple diseases and hence opens up the possibility of using an existing drug on other diseases as well as designing a single drug for multiple diseases.
Conflict of interest statement
Figures
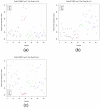
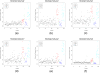
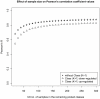

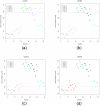
Similar articles
-
Tumor classification and marker gene prediction by feature selection and fuzzy c-means clustering using microarray data.BMC Bioinformatics. 2003 Dec 2;4:60. doi: 10.1186/1471-2105-4-60. BMC Bioinformatics. 2003. PMID: 14651757 Free PMC article.
-
Joint analysis of two microarray gene-expression data sets to select lung adenocarcinoma marker genes.BMC Bioinformatics. 2004 Jun 24;5:81. doi: 10.1186/1471-2105-5-81. BMC Bioinformatics. 2004. PMID: 15217521 Free PMC article.
-
Discovery of dominant and dormant genes from expression data using a novel generalization of SNR for multi-class problems.BMC Bioinformatics. 2008 Oct 9;9:425. doi: 10.1186/1471-2105-9-425. BMC Bioinformatics. 2008. PMID: 18842155 Free PMC article.
-
Cross-platform comparison and visualisation of gene expression data using co-inertia analysis.BMC Bioinformatics. 2003 Nov 21;4:59. doi: 10.1186/1471-2105-4-59. BMC Bioinformatics. 2003. PMID: 14633289 Free PMC article.
-
Signature Evaluation Tool (SET): a Java-based tool to evaluate and visualize the sample discrimination abilities of gene expression signatures.BMC Bioinformatics. 2008 Jan 28;9:58. doi: 10.1186/1471-2105-9-58. BMC Bioinformatics. 2008. PMID: 18221568 Free PMC article.
Cited by
-
Discovering monotonic stemness marker genes from time-series stem cell microarray data.BMC Genomics. 2015;16 Suppl 2(Suppl 2):S2. doi: 10.1186/1471-2164-16-S2-S2. Epub 2015 Jan 21. BMC Genomics. 2015. PMID: 25708300 Free PMC article.
-
Genetic algorithm based cancerous gene identification from microarray data using ensemble of filter methods.Med Biol Eng Comput. 2019 Jan;57(1):159-176. doi: 10.1007/s11517-018-1874-4. Epub 2018 Aug 1. Med Biol Eng Comput. 2019. PMID: 30069674
References
-
- Tsai YS, Lin CT, Tseng GC, Chung IF, Pal NR. Discovery of dominant and dormant genes from expression data using a novel generalization of SNR for multi-class problems. BMC Bioinformatics. 2008;9:425. doi: 10.1186/1471-2105-9-425. - DOI - PMC - PubMed
-
- Golub TR, Slonim DK, Tamayo P, Huard C, Gaasenbeek M, et al. Molecular classification of cancer: class discovery and class prediction by gene expression monitoring. Science. 1999;286:531–537. doi: 10.1126/science.286.5439.531. - DOI - PubMed
-
- Dudoit S, Fridlyand J, Speed TP. Comparison of discrimination methods for the classification of tumors using gene expression data. J Am Stat Assoc. 2002;97:77–87. doi: 10.1198/016214502753479248. - DOI
-
- Guyon I, Weston J, Barnhill S, Vapnik V. Gene selection for cancer classification using support vector machines. Mach Learn. 2002;46:389–422.
-
- Pal NR, Aguan K, Sharma A, Amari SI. Discovering biomarkers from gene expression data for predicting cancer subgroups using neural networks and relational fuzzy clustering. BMC Bioinformatics. 2007;8:5. doi: 10.1186/1471-2105-8-5. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

