Targeting several CAG expansion diseases by a single antisense oligonucleotide
- PMID: 21909428
- PMCID: PMC3164722
- DOI: 10.1371/journal.pone.0024308
Targeting several CAG expansion diseases by a single antisense oligonucleotide
Abstract
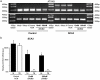
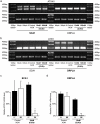
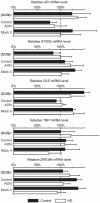
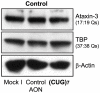
To date there are 9 known diseases caused by an expanded polyglutamine repeat, with the most prevalent being Huntington's disease. Huntington's disease is a progressive autosomal dominant neurodegenerative disorder for which currently no therapy is available. It is caused by a CAG repeat expansion in the HTT gene, which results in an expansion of a glutamine stretch at the N-terminal end of the huntingtin protein. This polyglutamine expansion plays a central role in the disease and results in the accumulation of cytoplasmic and nuclear aggregates. Here, we make use of modified 2'-O-methyl phosphorothioate (CUG)n triplet-repeat antisense oligonucleotides to effectively reduce mutant huntingtin transcript and protein levels in patient-derived Huntington's disease fibroblasts and lymphoblasts. The most effective antisense oligonucleotide, (CUG)(7), also reduced mutant ataxin-1 and ataxin-3 mRNA levels in spinocerebellar ataxia 1 and 3, respectively, and atrophin-1 in dentatorubral-pallidoluysian atrophy patient derived fibroblasts. This antisense oligonucleotide is not only a promising therapeutic tool to reduce mutant huntingtin levels in Huntington's disease but our results in spinocerebellar ataxia and dentatorubral-pallidoluysian atrophy cells suggest that this could also be applicable to other polyglutamine expansion disorders as well.
Conflict of interest statement
Figures



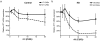
References
-
- Cummings CJ, Zoghbi HY. Fourteen and counting: unraveling trinucleotide repeat diseases. Hum Mol Genet. 2000;9:909–916. - PubMed
-
- Nakamura K, Jeong SY, Uchihara T, Anno M, Nagashima K, et al. SCA17, a novel autosomal dominant cerebellar ataxia caused by an expanded polyglutamine in TATA-binding protein. Hum Mol Genet. 2001;10:1441–1448. - PubMed
-
- Bauer PO, Nukina N. The pathogenic mechanisms of polyglutamine diseases and current therapeutic strategies. J Neurochem. 2009;110:1737–1765. - PubMed
-
- Mcneil SM, Novelletto A, Srinidhi J, Barnes G, Kornbluth I, et al. Reduced penetrance of the Huntington's disease mutation. Hum Mol Genet. 1997;6:775–779. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

