Phospholipase A2 enzymes: physical structure, biological function, disease implication, chemical inhibition, and therapeutic intervention
- PMID: 21910409
- PMCID: PMC3196595
- DOI: 10.1021/cr200085w
Phospholipase A2 enzymes: physical structure, biological function, disease implication, chemical inhibition, and therapeutic intervention
Figures
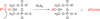
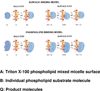






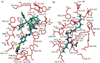




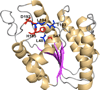
References
-
- Dennis EA. In: The Enzymes. Boyer P, editor. Vol. 16. New York: Academic Press; 1983.
-
- Davidson FF, Dennis EA. J. Mol. Evol. 1990;31:228. - PubMed
- Davidson FF, Dennis EA. In: Handbook of Natural Toxins. Tu AT, editor. Vol. 5. New York: Marcel Dekker; 1991.
-
- Seilhamer JJ, Pruzanski W, Vadas P, Plant S, Miller JA, Kloss J, Johnson LK. J. Biol. Chem. 1989;264:5335. - PubMed
-
- Kramer RM, Hession C, Johansen B, Hayes G, McGray P, Chow EP, Tizard R, Pepinsky RB. J. Biol. Chem. 1989;264:5768. - PubMed
-
- Clark JD, Lin LL, Kriz RW, Ramesha CS, Sultzman LA, Lin AY, Milona N, Knopf JL. Cell. 1991;65:1043. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

