A three-part signal governs differential processing of Gli1 and Gli3 proteins by the proteasome
- PMID: 21921029
- PMCID: PMC3234730
- DOI: 10.1074/jbc.M111.274993
A three-part signal governs differential processing of Gli1 and Gli3 proteins by the proteasome
Abstract
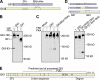
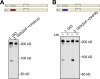
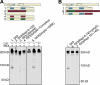
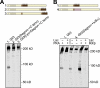
The Gli proteins are the transcriptional effectors of the mammalian Hedgehog signaling pathway. In an unusual mechanism, the proteasome partially degrades or processes Gli3 in the absence of Hedgehog pathway stimulation to create a Gli3 fragment that opposes the activity of the full-length protein. In contrast, Gli1 is not processed but degraded completely, despite considerable homology with Gli3. We found that these differences in processing can be described by defining a processing signal that is composed of three parts: the zinc finger domain, an adjacent linker sequence, and a degron. Gli3 processing is inhibited when any one component of the processing signal is disrupted. We show that the zinc fingers are required for processing only as a folded structure and that the location but not the identity of the processing degron is critical. Within the linker sequence, regions of low sequence complexity play a crucial role, but other sequence features are also important. Gli1 is not processed because two components of the processing signal, the linker sequence and the degron, are ineffective. These findings provide new insights into the molecular elements that regulate Gli protein processing by the proteasome.
Figures
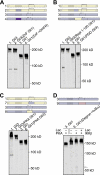

References
-
- Hooper J. E., Scott M. P. (2005) Nat. Rev. Mol. Cell Biol. 6, 306–317 - PubMed
-
- Kent D., Bush E. W., Hooper J. E. (2006) Development 133, 2001–2010 - PubMed
-
- Zhang Q., Zhang L., Wang B., Ou C. Y., Chien C. T., Jiang J. (2006) Dev. Cell 10, 719–729 - PubMed
-
- Aza-Blanc P., Ramírez-Weber F. A., Laget M. P., Schwartz C., Kornberg T. B. (1997) Cell 89, 1043–1053 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

