Crystal structure of a monomeric retroviral protease solved by protein folding game players
- PMID: 21926992
- PMCID: PMC3705907
- DOI: 10.1038/nsmb.2119
Crystal structure of a monomeric retroviral protease solved by protein folding game players
Erratum in
- Nat Struct Mol Biol. 2012 Mar;19(3):364
Abstract
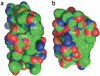
Following the failure of a wide range of attempts to solve the crystal structure of M-PMV retroviral protease by molecular replacement, we challenged players of the protein folding game Foldit to produce accurate models of the protein. Remarkably, Foldit players were able to generate models of sufficient quality for successful molecular replacement and subsequent structure determination. The refined structure provides new insights for the design of antiretroviral drugs.
Figures
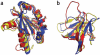

Comment in
-
Science and Culture: Putting a game face on biomedical research.Proc Natl Acad Sci U S A. 2016 Jun 14;113(24):6577-8. doi: 10.1073/pnas.1607585113. Proc Natl Acad Sci U S A. 2016. PMID: 27302944 Free PMC article. No abstract available.
References
-
- Rohl CA, Strauss CE, Misura KM, Baker D. Methods Enzymol. 2004;383:66–93. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

