Genetic evidence supporting the association of protease and protease inhibitor genes with inflammatory bowel disease: a systematic review
- PMID: 21931648
- PMCID: PMC3169567
- DOI: 10.1371/journal.pone.0024106
Genetic evidence supporting the association of protease and protease inhibitor genes with inflammatory bowel disease: a systematic review
Abstract
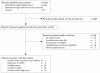
As part of the European research consortium IBDase, we addressed the role of proteases and protease inhibitors (P/PIs) in inflammatory bowel disease (IBD), characterized by chronic mucosal inflammation of the gastrointestinal tract, which affects 2.2 million people in Europe and 1.4 million people in North America. We systematically reviewed all published genetic studies on populations of European ancestry (67 studies on Crohn's disease [CD] and 37 studies on ulcerative colitis [UC]) to identify critical genomic regions associated with IBD. We developed a computer algorithm to map the 807 P/PI genes with exact genomic locations listed in the MEROPS database of peptidases onto these critical regions and to rank P/PI genes according to the accumulated evidence for their association with CD and UC. 82 P/PI genes (75 coding for proteases and 7 coding for protease inhibitors) were retained for CD based on the accumulated evidence. The cylindromatosis/turban tumor syndrome gene (CYLD) on chromosome 16 ranked highest, followed by acylaminoacyl-peptidase (APEH), dystroglycan (DAG1), macrophage-stimulating protein (MST1) and ubiquitin-specific peptidase 4 (USP4), all located on chromosome 3. For UC, 18 P/PI genes were retained (14 proteases and 4 protease inhibitors), with a considerably lower amount of accumulated evidence. The ranking of P/PI genes as established in this systematic review is currently used to guide validation studies of candidate P/PI genes, and their functional characterization in interdisciplinary mechanistic studies in vitro and in vivo as part of IBDase. The approach used here overcomes some of the problems encountered when subjectively selecting genes for further evaluation and could be applied to any complex disease and gene family.
Conflict of interest statement
Figures
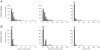


References
-
- Loftus EV, Jr, Sandborn WJ. Epidemiology of inflammatory bowel disease. Gastroenterol Clin North Am. 2002;31:1–20. - PubMed
-
- Loftus EV., Jr Clinical epidemiology of inflammatory bowel disease: Incidence, prevalence, and environmental influences. 2004;126:1504–1517. - PubMed
-
- Xavier RJ, Podolsky DK. Unravelling the pathogenesis of inflammatory bowel disease. 2007;448:427–434. - PubMed
-
- Hugot JP, Chamaillard M, Zouali H, Lesage S, Cezard JP, et al. Association of NOD2 leucine-rich repeat variants with susceptibility to Crohn's disease. 2001;411:599–603. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

