Accuracy of genomic selection in simulated populations mimicking the extent of linkage disequilibrium in beef cattle
- PMID: 21933416
- PMCID: PMC3224120
- DOI: 10.1186/1471-2156-12-80
Accuracy of genomic selection in simulated populations mimicking the extent of linkage disequilibrium in beef cattle
Abstract
Background: The success of genomic selection depends mainly on the extent of linkage disequilibrium (LD) between markers and quantitative trait loci (QTL), the number of animals in the training set (TS) and the heritability (h2) of the trait. The extent of LD depends on the genetic structure of the population and the density of markers. The aim of this study was to calculate accuracy of direct genomic estimated breeding values (DGEBV) using best linear unbiased genomic prediction (GBLUP) for different marker densities, heritabilities and sizes of the TS in simulated populations that mimicked previously reported extent and pattern of LD in beef cattle.
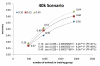
Results: The accuracy of DGEBV increased significantly (p < 0.05) with the increase in the number of bulls in the TS (480, 960 or 1920), trait h2 (0.10, 0.25 or 0.40) and marker densities (40 k or 800 k). Increasing the number of animals in the TS by 4-fold and using their phenotypes to estimate marker effects was not sufficient to maintain or increase the accuracy of DGEBV obtained using estimated breeding values (EBVs) when the trait h2 was lower than 0.40 for both marker densities. Comparing to expected accuracies of parent average (PA), the gains by using DGEBV would be of 27%, 13% and 10% for trait h2 equal to 0.10, 0.25 and 0.40, respectively, considering the scenario with 40 k markers and 1920 bulls in TS.
Conclusions: As reported in dairy cattle, the size of the TS and the extent of LD have major impact on the accuracy of DGEBV. Based on the findings of this simulation study, large TS, as well as dense marker panels, aiming to increase the level of LD between markers and QTL, will likely be needed in beef cattle for successful implementation of genomic selection.
Figures
Similar articles
-
Genomic prediction ability for beef fatty acid profile in Nelore cattle using different pseudo-phenotypes.J Appl Genet. 2018 Nov;59(4):493-501. doi: 10.1007/s13353-018-0470-5. Epub 2018 Sep 24. J Appl Genet. 2018. PMID: 30251238
-
Empirical and deterministic accuracies of across-population genomic prediction.Genet Sel Evol. 2015 Feb 6;47(1):5. doi: 10.1186/s12711-014-0086-0. Genet Sel Evol. 2015. PMID: 25885467 Free PMC article.
-
Different models of genetic variation and their effect on genomic evaluation.Genet Sel Evol. 2011 May 17;43(1):18. doi: 10.1186/1297-9686-43-18. Genet Sel Evol. 2011. PMID: 21575265 Free PMC article.
-
Invited review: Genomic selection in dairy cattle: progress and challenges.J Dairy Sci. 2009 Feb;92(2):433-43. doi: 10.3168/jds.2008-1646. J Dairy Sci. 2009. PMID: 19164653 Review.
-
Genomic selection.J Anim Breed Genet. 2007 Dec;124(6):323-30. doi: 10.1111/j.1439-0388.2007.00702.x. J Anim Breed Genet. 2007. PMID: 18076469 Review.
Cited by
-
Weighted GBLUP in Simulated Beef Cattle Populations: Impact of Reference Population, Marker Density, and Heritability.Animals (Basel). 2025 Apr 12;15(8):1118. doi: 10.3390/ani15081118. Animals (Basel). 2025. PMID: 40281952 Free PMC article.
-
Accuracy of genome-wide imputation in Braford and Hereford beef cattle.BMC Genet. 2014 Dec 29;15:157. doi: 10.1186/s12863-014-0157-9. BMC Genet. 2014. PMID: 25543517 Free PMC article.
-
Combining Genome-Wide Information with a Functional Structural Plant Model to Simulate 1-Year-Old Apple Tree Architecture.Front Plant Sci. 2017 Jan 12;7:2065. doi: 10.3389/fpls.2016.02065. eCollection 2016. Front Plant Sci. 2017. PMID: 28127302 Free PMC article.
-
Non-additive association analysis using proxy phenotypes identifies novel cattle syndromes.Nat Genet. 2021 Jul;53(7):949-954. doi: 10.1038/s41588-021-00872-5. Epub 2021 May 27. Nat Genet. 2021. PMID: 34045765
-
Multi-population genomic prediction using a multi-task Bayesian learning model.BMC Genet. 2014 May 3;15:53. doi: 10.1186/1471-2156-15-53. BMC Genet. 2014. PMID: 24884927 Free PMC article.
References
-
- Schenkel FS, Sargolzaei M, Kistemaker G, Jansen GB, Sullivan P, Van Doormaal BJ, VanRaden PM, Wiggans GR. Reliability of genomic evaluation of Holstein cattle in Canada. Proccedings of Interbull International Workshop - genomic information in genetic evaluations Uppsala, Sweden. 2009. http://www-interbull.slu.se/bulletins/framesida-pub.htm
-
- Taylor JF, Decker JE, Kim J, McClure MC, Mckay SD, Rolf MM, Taxis T, Vasco D, Schnabel RD. Prospects for genomic selection in beef cattle. Proceedings of the 41th Beef Improvement Federation Annual Research Symposium: Sacramento, USA. 2009.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

