The nucleolar protein Nop19p interacts preferentially with Utp25p and Dhr2p and is essential for the production of the 40S ribosomal subunit in Saccharomyces cerevisiae
- PMID: 21941128
- PMCID: PMC3256425
- DOI: 10.4161/rna.8.6.17699
The nucleolar protein Nop19p interacts preferentially with Utp25p and Dhr2p and is essential for the production of the 40S ribosomal subunit in Saccharomyces cerevisiae
Abstract
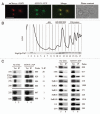
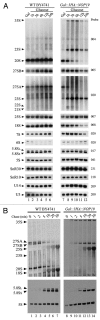
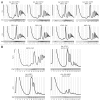
In eukaryotes, ribosome biogenesis is a process of major interest that requires more than 200 factors acting coordinately in time and space. Using genetic and proteomic studies, most of the components have now been identified. Based on its nucleolar localization, we characterized the protein encoded by the open reading frame YGR251W, we renamed Nop19p as playing an essential role in ribosome biogenesis. Depletion of the Nop19p in yeast impairs pre-rRNA processing at sites A₀, A₁ and A₂, leading to a strong decrease in 18S rRNA and 40S subunit levels. Nop19p is a component of 90S preribosomes which assembly is believed to result from stepwise incorporation of UTP modules. We show that Nop19p depletion does not impair the incorporation of UTP subcomplexes on preribosomes and conversely that depletion of UTP subcomplexes does not affect Nop19p recruitment on 90S preribosomes. TAP experiments under stringent conditions revealed that Nop19p interacts preferentially with the DEAH-box RNA helicase Dhr2p and Utp25p, both required for A 0, A 1 and A 2 cleavages. Nop19p appeared essential for the incorporation of Utp25p in preribosomes. In addition, our results suggest that in absence of Nop19p, Dhr2p remains trapped within aberrant preribosomes.
Figures




Similar articles
-
Evolutionarily conserved function of RRP36 in early cleavages of the pre-rRNA and production of the 40S ribosomal subunit.Mol Cell Biol. 2010 Mar;30(5):1130-44. doi: 10.1128/MCB.00999-09. Epub 2009 Dec 28. Mol Cell Biol. 2010. PMID: 20038530 Free PMC article.
-
hUTP24 is essential for processing of the human rRNA precursor at site A1, but not at site A0.RNA Biol. 2015;12(9):1010-29. doi: 10.1080/15476286.2015.1073437. RNA Biol. 2015. PMID: 26237581 Free PMC article.
-
Utp25p, a nucleolar Saccharomyces cerevisiae protein, interacts with U3 snoRNP subunits and affects processing of the 35S pre-rRNA.FEBS J. 2010 Jul;277(13):2838-52. doi: 10.1111/j.1742-4658.2010.07701.x. Epub 2010 May 27. FEBS J. 2010. PMID: 20528918
-
Inside the 40S ribosome assembly machinery.Curr Opin Chem Biol. 2011 Oct;15(5):657-63. doi: 10.1016/j.cbpa.2011.07.023. Epub 2011 Aug 20. Curr Opin Chem Biol. 2011. PMID: 21862385 Free PMC article. Review.
-
Ribosome biogenesis in the yeast Saccharomyces cerevisiae.Genetics. 2013 Nov;195(3):643-81. doi: 10.1534/genetics.113.153197. Genetics. 2013. PMID: 24190922 Free PMC article. Review.
Cited by
-
RNA folding and functions of RNA helicases in ribosome biogenesis.RNA Biol. 2022 Jan;19(1):781-810. doi: 10.1080/15476286.2022.2079890. RNA Biol. 2022. PMID: 35678541 Free PMC article. Review.
-
Ribosome biogenesis factors-from names to functions.EMBO J. 2023 Apr 3;42(7):e112699. doi: 10.15252/embj.2022112699. Epub 2023 Feb 10. EMBO J. 2023. PMID: 36762427 Free PMC article. Review.
-
DExD/H-box RNA helicases in ribosome biogenesis.RNA Biol. 2013 Jan;10(1):4-18. doi: 10.4161/rna.21879. Epub 2012 Aug 24. RNA Biol. 2013. PMID: 22922795 Free PMC article. Review.
-
Turnover of aberrant pre-40S pre-ribosomal particles is initiated by a novel endonucleolytic decay pathway.Nucleic Acids Res. 2018 May 18;46(9):4699-4714. doi: 10.1093/nar/gky116. Nucleic Acids Res. 2018. PMID: 29481617 Free PMC article.
References
-
- Venema J, Tollervey D. Ribosome synthesis in Saccharomyces cerevisiae. Ann Rev Gen. 1999;33:261–311. - PubMed
-
- Lafontaine DL, Tollervey D. The function and synthesis of ribosomes. Nat Rev Mol Cell Biol. 2001;2:514–520. - PubMed
-
- de la Cruz J, Kressler D, Linder P. Ribosomal subunit assembly. In: Olson MOJ, editor. The Nucleolus. KluwerAcademic/Plenum Publishers; 2003. pp. 262–290.
-
- Fromont-Racine M, Senger B, Saveanu C, Fasiolo F. Ribosome assembly in eukaryotes. Gene. 2003;313:17–42. - PubMed
-
- Dez C, Tollervey D. Ribosome synthesis meets the cell cycle. Curr Opin Microbiol. 2004;7:631–637. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous
