N-acetylglucosamine-6-phosphate deacetylase (NagA) of Listeria monocytogenes EGD, an essential enzyme for the metabolism and recycling of amino sugars
- PMID: 21947170
- PMCID: PMC3304070
- DOI: 10.1007/s00203-011-0752-3
N-acetylglucosamine-6-phosphate deacetylase (NagA) of Listeria monocytogenes EGD, an essential enzyme for the metabolism and recycling of amino sugars
Abstract
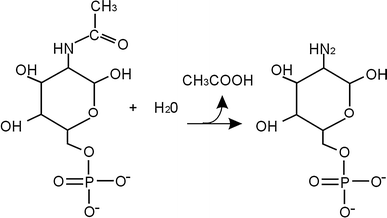
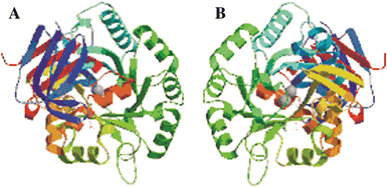
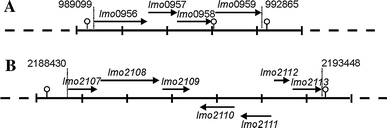
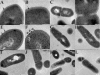
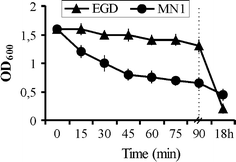
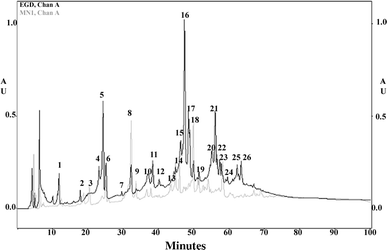
The main aim of our study was to determine the physiological function of NagA enzyme in the Listeria monocytogenes cell. The primary structure of the murein of L. monocytogenes is very similar to that of Escherichia coli, the main differences being amidation of diaminopimelic acid and partial de-N-acetylation of glucosamine residues. NagA is needed for the deacetylation of N-acetyl-glucosamine-6 phosphate to glucosamine-6 phosphate and acetate. Analysis of the L. monocytogenes genome reveals the presence of two proteins with NagA domain, Lmo0956 and Lmo2108, which are cytoplasmic putative proteins. We introduced independent mutations into the structural genes for the two proteins. In-depth characterization of one of these mutants, MN1, deficient in protein Lmo0956 revealed strikingly altered cell morphology, strongly reduced cell wall murein content and decreased sensitivity to cell wall hydrolase, mutanolysin and peptide antibiotic, colistin. The gene products of operon 150, consisting of three genes: lmo0956, lmo0957, and lmo0958, are necessary for the cytosolic steps of the amino-sugar-recycling pathway. The cytoplasmic de-N-acetylase Lmo0956 of L. monocytogenes is required for cell wall peptidoglycan and teichoic acid biosynthesis and is also essential for bacterial cell growth, cell division, and sensitivity to cell wall hydrolases and peptide antibiotics.
Figures


Similar articles
-
Inactivation of the wall-associated de-N-acetylase (PgdA) of Listeria monocytogenes results in greater susceptibility of the cells to induced autolysis.J Microbiol Biotechnol. 2009 Sep;19(9):932-45. doi: 10.4014/jmb.0810.557. J Microbiol Biotechnol. 2009. PMID: 19809250
-
Cloning and characterization of the N-acetylglucosamine operon of Escherichia coli.Biochem Cell Biol. 1990 Jan;68(1):123-37. doi: 10.1139/o90-017. Biochem Cell Biol. 1990. PMID: 2190615
-
The gate controlling cell wall synthesis in Staphylococcus aureus.Mol Microbiol. 2004 Aug;53(4):1221-31. doi: 10.1111/j.1365-2958.2004.04200.x. Mol Microbiol. 2004. PMID: 15306023
-
Analysis of the peptidoglycan hydrolases of Listeria monocytogenes: multiple enzymes with multiple functions.Pol J Microbiol. 2004;53 Suppl:29-34. Pol J Microbiol. 2004. PMID: 15787194 Review.
-
The Cell Wall of Streptococcus pneumoniae.Microbiol Spectr. 2019 May;7(3):10.1128/microbiolspec.gpp3-0018-2018. doi: 10.1128/microbiolspec.GPP3-0018-2018. Microbiol Spectr. 2019. PMID: 31172911 Free PMC article. Review.
Cited by
-
The Analysis of Field Strains Isolated From Food, Animal and Clinical Sources Uncovers Natural Mutations in Listeria monocytogenes Nisin Resistance Genes.Front Microbiol. 2020 Oct 6;11:549531. doi: 10.3389/fmicb.2020.549531. eCollection 2020. Front Microbiol. 2020. PMID: 33123101 Free PMC article.
-
Chitin Attenuates Expression of Listeria monocytogenes Virulence Genes in vitro.Front Microbiol. 2020 Dec 3;11:588906. doi: 10.3389/fmicb.2020.588906. eCollection 2020. Front Microbiol. 2020. PMID: 33343529 Free PMC article.
-
Investigating endophytic fungi of Calotropis procera for novel bioactive compounds: molecular docking and bioactivity insights.Microb Cell Fact. 2025 May 8;24(1):101. doi: 10.1186/s12934-025-02710-3. Microb Cell Fact. 2025. PMID: 40340934 Free PMC article.
-
Analysis of temporal gene regulation of Listeria monocytogenes revealed distinct regulatory response modes after exposure to high pressure processing.BMC Genomics. 2021 Apr 14;22(1):266. doi: 10.1186/s12864-021-07461-0. BMC Genomics. 2021. PMID: 33853520 Free PMC article.
-
Delving into the Characteristic Features of "Menace" Mycobacterium tuberculosis Homologs: A Structural Dynamics and Proteomics Perspectives.Protein J. 2020 Apr;39(2):118-132. doi: 10.1007/s10930-020-09890-4. Protein J. 2020. PMID: 32162114
References
-
- Ames BN. Assay of inorganic phosphate, total phosphate, and phosphatases. Methods Enzymol. 1966;8:115–118. doi: 10.1016/0076-6879(66)08014-5. - DOI
-
- Arunkumar K, Manoj N (2010) Molecular evolution of the N-acetylglucosamine-6-phosphate deacetylase gene. In: ISB ‘10 Proceedings of the international symposium on biocomputing. ACM, New York. doi: 10.1145/1722024.1722076
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

