The S100A10-annexin A2 complex provides a novel asymmetric platform for membrane repair
- PMID: 21949189
- PMCID: PMC3220529
- DOI: 10.1074/jbc.M111.244038
The S100A10-annexin A2 complex provides a novel asymmetric platform for membrane repair
Erratum in
- J Biol Chem. 2011 Dec 16;286(50):43588
Abstract
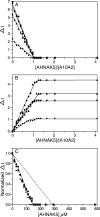
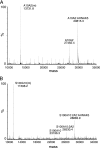
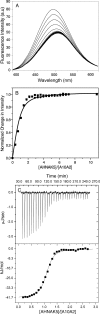
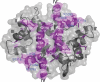
Membrane repair is mediated by multiprotein complexes, such as that formed between the dimeric EF-hand protein S100A10, the calcium- and phospholipid-binding protein annexin A2, the enlargeosome protein AHNAK, and members of the transmembrane ferlin family. Although interactions between these proteins have been shown, little is known about their structural arrangement and mechanisms of formation. In this work, we used a non-covalent complex between S100A10 and the N terminus of annexin A2 (residues 1-15) and a designed hybrid protein (A10A2), where S100A10 is linked in tandem to the N-terminal region of annexin A2, to explore the binding region, stoichiometry, and affinity with a synthetic peptide from the C terminus of AHNAK. Using multiple biophysical methods, we identified a novel asymmetric arrangement between a single AHNAK peptide and the A10A2 dimer. The AHNAK peptide was shown to require the annexin A2 N terminus, indicating that the AHNAK binding site comprises regions on both S100A10 and annexin proteins. NMR spectroscopy was used to show that the AHNAK binding surface comprised residues from helix IV in S100A10 and the C-terminal portion from the annexin A2 peptide. This novel surface maps to the exposed side of helices IV and IV' of the S100 dimeric structure, a region not identified in any previous S100 target protein structures. The results provide the first structural details of the ternary S100A10 protein complex required for membrane repair.
Figures


Similar articles
-
Structure of an asymmetric ternary protein complex provides insight for membrane interaction.Structure. 2012 Oct 10;20(10):1737-45. doi: 10.1016/j.str.2012.08.004. Epub 2012 Aug 30. Structure. 2012. PMID: 22940583
-
Structure of a C-terminal AHNAK peptide in a 1:2:2 complex with S100A10 and an acetylated N-terminal peptide of annexin A2.Acta Crystallogr D Biol Crystallogr. 2013 Jan;69(Pt 1):92-104. doi: 10.1107/S0907444912043429. Epub 2012 Dec 20. Acta Crystallogr D Biol Crystallogr. 2013. PMID: 23275167 Free PMC article.
-
Design of high-affinity S100-target hybrid proteins.Protein Sci. 2009 Dec;18(12):2528-36. doi: 10.1002/pro.267. Protein Sci. 2009. PMID: 19827097 Free PMC article.
-
The Annexin A2/S100A10 Complex: The Mutualistic Symbiosis of Two Distinct Proteins.Biomolecules. 2021 Dec 9;11(12):1849. doi: 10.3390/biom11121849. Biomolecules. 2021. PMID: 34944495 Free PMC article. Review.
-
Unique S100 target protein interactions.Gen Physiol Biophys. 2009;28 Spec No Focus:F39-46. Gen Physiol Biophys. 2009. PMID: 20093725 Review.
Cited by
-
Functions of S100 proteins.Curr Mol Med. 2013 Jan;13(1):24-57. Curr Mol Med. 2013. PMID: 22834835 Free PMC article. Review.
-
Connexin 30 Deficiency Attenuates Chronic but Not Acute Phases of Experimental Autoimmune Encephalomyelitis Through Induction of Neuroprotective Microglia.Front Immunol. 2018 Nov 7;9:2588. doi: 10.3389/fimmu.2018.02588. eCollection 2018. Front Immunol. 2018. PMID: 30464764 Free PMC article.
-
CPT1A-mediated succinylation of S100A10 increases human gastric cancer invasion.J Cell Mol Med. 2019 Jan;23(1):293-305. doi: 10.1111/jcmm.13920. Epub 2018 Nov 5. J Cell Mol Med. 2019. PMID: 30394687 Free PMC article.
-
Role of calcium-sensor proteins in cell membrane repair.Biosci Rep. 2023 Feb 27;43(2):BSR20220765. doi: 10.1042/BSR20220765. Biosci Rep. 2023. PMID: 36728029 Free PMC article. Review.
-
Silencing AHNAK promotes nasopharyngeal carcinoma progression by upregulating the ANXA2 protein.Cell Oncol (Dordr). 2024 Jun;47(3):833-850. doi: 10.1007/s13402-023-00898-3. Epub 2023 Nov 14. Cell Oncol (Dordr). 2024. PMID: 37962808
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

