Hormone replacement therapy dependent changes in breast cancer-related gene expression in breast tissue of healthy postmenopausal women
- PMID: 21956102
- PMCID: PMC5528323
- DOI: 10.1016/j.molonc.2011.09.003
Hormone replacement therapy dependent changes in breast cancer-related gene expression in breast tissue of healthy postmenopausal women
Abstract
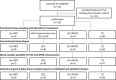
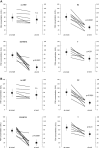
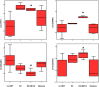
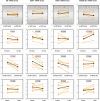
Risk assessment of future breast cancer risk through exposure to sex steroids currently relies on clinical scorings such as mammographic density. Knowledge about the gene expression patterns in existing breast cancer tumors may be used to identify risk factors in the breast tissue of women still free of cancer. The differential effects of estradiol, estradiol together with gestagens, or tibolone on breast cancer-related gene expression in normal breast tissue samples taken from postmenopausal women may be used to identify gene expression profiles associated with a higher breast cancer risk. Breast tissue samples were taken from 33 healthy postmenopausal women both before and after a six month treatment with either 2mg micronized estradiol [E2], 2mg micronized estradiol and 1mg norethisterone acetate [E2+NETA], 2.5mg tibolone [T] or [no HRT]. Except for [E2], which was only given to women after hysterectomy, the allocation to each of the three groups was randomized. The expression of 102 mRNAs and 46 microRNAs putatively involved in breast cancer was prospectively determined in the biopsies of 6 women receiving [no HRT], 5 women receiving [E2], 5 women receiving [E2+NETA], and 6 receiving [T]. Using epithelial and endothelial markers genes, non-representative biopsies from 11 women were eliminated. Treatment of postmenopausal women with [E2+NETA] resulted in the highest number of differentially (p<0.05) regulated genes (16.2%) compared to baseline, followed by [E2] (10.1%) and [T] (4.7%). Among genes that were significantly down-regulated by [E2+NETA] ranked estrogen-receptor-1 (ESR1, p=0.019) and androgen receptor (AR, p=0.019), whereas CYP1B1, a gene encoding an estrogen-metabolizing enzyme, was significantly up-regulated (p=0.016). Mammary cells triggered by [E2+NETA] and [E2] adjust for steroidogenic up-regulation through down-regulation of the estrogen-receptor pathway. In this prospective study, prolonged administration of [E2+NETA] and to a lesser extent of [E2] but not [T] were associated in otherwise healthy breast tissue with a change in the expression of genes putatively involved in breast cancer. Our data suggest that normal mammary cells triggered by [E2+NETA] adjust for steroidogenic up-regulation through down-regulation of the estrogen-receptor pathway. This feasibility study provides the basis for whole genome analyses to identify novel markers involved in increased breast cancer risk.
Copyright © 2011 Federation of European Biochemical Societies. Published by Elsevier B.V. All rights reserved.
Figures

Similar articles
-
Economic impact of tibolone compared with continuous-combined hormone replacement therapy. In the management of postmenopausal women with climacteric symptoms in the UK.Pharmacoeconomics. 2000 Nov;18(5):477-86. doi: 10.2165/00019053-200018050-00007. Pharmacoeconomics. 2000. PMID: 11151401
-
Effects of tibolone and continuous combined hormone replacement therapy on mammographic breast density.Am J Obstet Gynecol. 2002 Apr;186(4):717-22. doi: 10.1067/mob.2002.121896. Am J Obstet Gynecol. 2002. PMID: 11967497 Clinical Trial.
-
A microarray study on the effect of four hormone therapy regimens on gene transcription in whole blood from healthy postmenopausal women.Thromb Res. 2012 Jul;130(1):45-51. doi: 10.1016/j.thromres.2011.12.009. Epub 2012 Jan 2. Thromb Res. 2012. PMID: 22217510 Clinical Trial.
-
[Tibolone].Presse Med. 2002 Sep 7;31(28):1314-22. Presse Med. 2002. PMID: 12355994 Review. French.
-
Effects of tibolone on the breast of postmenopausal women.Taiwan J Obstet Gynecol. 2007 Jun;46(2):121-6. doi: 10.1016/S1028-4559(07)60005-9. Taiwan J Obstet Gynecol. 2007. PMID: 17638619 Review.
Cited by
-
Effects of Estradiol/Micronized Progesterone vs. Conjugated Equine Estrogens/Medroxyprogesterone Acetate on Breast Cancer Gene Expression in Healthy Postmenopausal Women.Int J Mol Sci. 2023 Feb 18;24(4):4123. doi: 10.3390/ijms24044123. Int J Mol Sci. 2023. PMID: 36835533 Free PMC article. Clinical Trial.
-
Hormone therapy use and breast tissue DNA methylation: analysis of epigenome wide data from the normal breast study.Epigenetics. 2019 Feb;14(2):146-157. doi: 10.1080/15592294.2019.1580111. Epub 2019 Mar 1. Epigenetics. 2019. PMID: 30821641 Free PMC article.
-
Associations of breast cancer related exposures and gene expression profiles in normal breast tissue-The Norwegian Women and Cancer normal breast tissue study.Cancer Rep (Hoboken). 2023 Apr;6(4):e1777. doi: 10.1002/cnr2.1777. Epub 2023 Jan 8. Cancer Rep (Hoboken). 2023. PMID: 36617746 Free PMC article.
-
Estrogen and androgen-converting enzymes 17β-hydroxysteroid dehydrogenase and their involvement in cancer: with a special focus on 17β-hydroxysteroid dehydrogenase type 1, 2, and breast cancer.Oncotarget. 2017 May 2;8(18):30552-30562. doi: 10.18632/oncotarget.15547. Oncotarget. 2017. PMID: 28430630 Free PMC article. Review.
References
-
- Asselin-Labat, M.L. , Vaillant, F. , Sheridan, J.M. , Pal, B. , Wu, D. , Simpson, E.R. , Yasuda, H. , Smyth, G.K. , Martin, T.J. , Lindeman, G.F. , Visvader, J.E. , 2010. Control of mammary stem cell function by steroid hormone signalling. Nature. 465, 798–802. - PubMed
-
- Banks, E. , Reeves, G. , Beral, V. , Bull, D. , Crossley, B. , Simmonds, M. , Hilton, E. , Bailey, S. , Barrett, N. , Briers, P. , 2006. Hormone replacement therapy and false positive recall in the Million Women Study: patterns of use, hormonal constituents and consistency of effect. Breast Cancer Res.. 8, R8 - PMC - PubMed
-
- Benjamini, Y. , Hochberg, Y. , 1995. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. Roy. Stat. Soc., Ser. B. 57, 289–300.
-
- Beral, V. , Million Women Study Collaborators, 2003. Breast cancer and hormone-replacement therapy in the Million Women Study. Lancet. 362, 419–427. - PubMed
-
- Byrne, C. , Schairer, C. , Wolfe, J. , Parekh, N. , Salane, M. , Brinton, L.A. , Hoover, R. , Haile, R. , 1995. Mammographic features and breast cancer risk: effects with time, age, and menopause status. J. Natl. Cancer Inst.. 87, 1622–1629. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

