Structural correlates of selectivity and inactivation in potassium channels
- PMID: 21958666
- PMCID: PMC3253935
- DOI: 10.1016/j.bbamem.2011.09.007
Structural correlates of selectivity and inactivation in potassium channels
Abstract
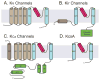
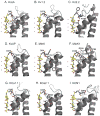
Potassium channels are involved in a tremendously diverse range of physiological applications requiring distinctly different functional properties. Not surprisingly, the amino acid sequences for these proteins are diverse as well, except for the region that has been ordained the "selectivity filter". The goal of this review is to examine our current understanding of the role of the selectivity filter and regions adjacent to it in specifying selectivity as well as its role in gating/inactivation and possible mechanisms by which these processes are coupled. Our working hypothesis is that an amino acid network behind the filter modulates selectivity in channels with the same signature sequence while at the same time affecting channel inactivation properties. This article is part of a Special Issue entitled: Membrane protein structure and function.
Copyright © 2011 Elsevier B.V. All rights reserved.
Figures

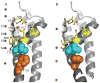
References
-
- Yu FH, Catterall WA. The VGL-chanome: a protein superfamily specialized for electrical signaling and ionic homeostasis. Sci STKE. 2004;2004:re15. - PubMed
-
- Yellen G. The voltage-gated potassium channels and their relatives. Nature. 2002;419:35–42. - PubMed
-
- Hibino H, Inanobe A, Furutani K, Murakami S, Findlay I, Kurachi Y. Inwardly rectifying potassium channels: their structure, function, and physiological roles. Physiol Rev. 2010;90:291–366. - PubMed
-
- Xia XM, Zeng X, Lingle CJ. Multiple regulatory sites in large-conductance calcium-activated potassium channels. Nature. 2002;418:880–884. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

