Integrating autoimmune risk loci with gene-expression data identifies specific pathogenic immune cell subsets
- PMID: 21963258
- PMCID: PMC3188838
- DOI: 10.1016/j.ajhg.2011.09.002
Integrating autoimmune risk loci with gene-expression data identifies specific pathogenic immune cell subsets
Erratum in
- Am J Hum Genet. 2011 Nov 11;89(5):682
Abstract
Although genome-wide association studies have implicated many individual loci in complex diseases, identifying the exact causal alleles and the cell types within which they act remains greatly challenging. To ultimately understand disease mechanism, researchers must carefully conceive functional studies in relevant pathogenic cell types to demonstrate the cellular impact of disease-associated genetic variants. This challenge is highlighted in autoimmune diseases, such as rheumatoid arthritis, where any of a broad range of immunological cell types might potentially be impacted by genetic variation to cause disease. To this end, we developed a statistical approach to identify potentially pathogenic cell types in autoimmune diseases by using a gene-expression data set of 223 murine-sorted immune cells from the Immunological Genome Consortium. We found enrichment of transitional B cell genes in systemic lupus erythematosus (p = 5.9 × 10(-6)) and epithelial-associated stimulated dendritic cell genes in Crohn disease (p = 1.6 × 10(-5)). Finally, we demonstrated enrichment of CD4+ effector memory T cell genes within rheumatoid arthritis loci (p < 10(-6)). To further validate the role of CD4+ effector memory T cells within rheumatoid arthritis, we identified 436 loci that were not yet known to be associated with the disease but that had a statistically suggestive association in a recent genome-wide association study (GWAS) meta-analysis (p(GWAS) < 0.001). Even among these putative loci, we noted a significant enrichment for genes specifically expressed in CD4+ effector memory T cells (p = 1.25 × 10(-4)). These cell types are primary candidates for future functional studies to reveal the role of risk alleles in autoimmunity. Our approach has application in other phenotypes, outside of autoimmunity, where many loci have been discovered and high-quality cell-type-specific gene expression is available.
Copyright © 2011 The American Society of Human Genetics. Published by Elsevier Inc. All rights reserved.
Figures

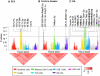

References
-
- Barrett J.C., Clayton D.G., Concannon P., Akolkar B., Cooper J.D., Erlich H.A., Julier C., Morahan G., Nerup J., Nierras C., Type 1 Diabetes Genetics Consortium Genome-wide association study and meta-analysis find that over 40 loci affect risk of type 1 diabetes. Nat. Genet. 2009;41:703–707. - PMC - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials

