TFIIH is an elongation factor of RNA polymerase I
- PMID: 21965540
- PMCID: PMC3258137
- DOI: 10.1093/nar/gkr746
TFIIH is an elongation factor of RNA polymerase I
Abstract
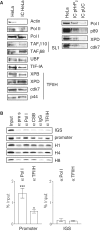
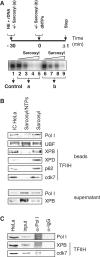
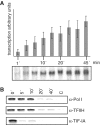
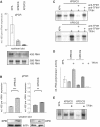
TFIIH is a multisubunit factor essential for transcription initiation and promoter escape of RNA polymerase II and for the opening of damaged DNA double strands in nucleotide excision repair (NER). In this study, we have analyzed at which step of the transcription cycle TFIIH is essential for transcription by RNA polymerase I. We demonstrate that TFIIH associates with the rDNA promoter and gene-internal sequences and leaves the rDNA promoter in a complex with RNA polymerase I after start of transcription. Moreover, mutations in the TFIIH subunits XPB and XPD found in Cockayne syndrome impair the interaction of TFIIH with the rDNA, but do not influence initiation complex formation or promoter escape of RNA polymerase I, but preclude the productivity of the enzyme by reducing transcription elongation in vivo and in vitro. Our results implicate that reduced RNA polymerase I transcription elongation and ribosomal stress could be one factor contributing to the Cockayne syndrome phenotype.
Figures


References
-
- Comai L, Tanese N, Tjian R. The TATA-binding protein and associated factors are integral components of the RNA polymerase I transcription factor, SL1. Cell. 1992;68:965–976. - PubMed
-
- Lobo SM, Tanaka M, Sullivan ML, Hernandez N. A TBP complex essential for transcription from TATA-less but not TATA-containing RNA polymerase III promoters is part of the TFIIIB fraction. Cell. 1992;71:1029–1040. - PubMed
-
- Iben S, Tschochner H, Bier M, Hoogstraten D, Hozak P, Egly JM, Grummt I. TFIIH plays an essential role in RNA polymerase I transcription. Cell. 2002;109:297–306. - PubMed
-
- Bradsher J, Auriol J, Proietti de Santis L, Iben S, Vonesch JL, Grummt I, Egly JM. CSB is a component of RNA pol I transcription. Mol. Cell. 2002;10:819–829. - PubMed
-
- Hoogstraten D, Nigg AL, Heath H, Mullenders LH, van Driel R, Hoeijmakers JH, Vermeulen W, Houtsmuller AB. Rapid switching of TFIIH between RNA polymerase I and II transcription and DNA repair in vivo. Mol. Cell. 2002;10:1163–1174. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

