Impact of the N-terminal secretor domain on YopD translocator function in Yersinia pseudotuberculosis type III secretion
- PMID: 21965570
- PMCID: PMC3232875
- DOI: 10.1128/JB.00210-11
Impact of the N-terminal secretor domain on YopD translocator function in Yersinia pseudotuberculosis type III secretion
Abstract
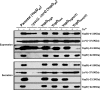
Type III secretion systems (T3SSs) secrete needle components, pore-forming translocators, and the translocated effectors. In part, effector recognition by a T3SS involves their N-terminal amino acids and their 5' mRNA. To investigate whether similar molecular constraints influence translocator secretion, we scrutinized this region within YopD from Yersinia pseudotuberculosis. Mutations in the 5' end of yopD that resulted in specific disruption of the mRNA sequence did not affect YopD secretion. On the other hand, a few mutations affecting the protein sequence reduced secretion. Translational reporter fusions identified the first five codons as a minimal N-terminal secretion signal and also indicated that the YopD N terminus might be important for yopD translation control. Hybrid proteins in which the N terminus of YopD was exchanged with the equivalent region of the YopE effector or the YopB translocator were also constructed. While the in vitro secretion profile was unaltered, these modified bacteria were all compromised with respect to T3SS activity in the presence of immune cells. Thus, the YopD N terminus does harbor a secretion signal that may also incorporate mechanisms of yopD translation control. This signal tolerates a high degree of variation while still maintaining secretion competence suggestive of inherent structural peculiarities that make it distinct from secretion signals of other T3SS substrates.
Figures









References
-
- Aili M., Isaksson E. L., Hallberg B., Wolf-Watz H., Rosqvist R. 2006. Functional analysis of the YopE GTPase-activating protein (GAP) activity of Yersinia pseudotuberculosis. Cell Microbiol. 8:1020–1033 - PubMed
-
- Akeda Y., Galan J. E. 2005. Chaperone release and unfolding of substrates in type III secretion. Nature 437:911–915 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

