Catalytic mechanism investigation of lysine-specific demethylase 1 (LSD1): a computational study
- PMID: 21984927
- PMCID: PMC3184146
- DOI: 10.1371/journal.pone.0025444
Catalytic mechanism investigation of lysine-specific demethylase 1 (LSD1): a computational study
Abstract
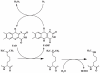
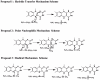
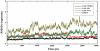
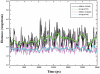
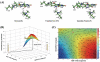
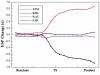
Lysine-specific demethylase 1 (LSD1), the first identified histone demethylase, is a flavin-dependent amine oxidase which specifically demethylates mono- or dimethylated H3K4 and H3K9 via a redox process. It participates in a broad spectrum of biological processes and is of high importance in cell proliferation, adipogenesis, spermatogenesis, chromosome segregation and embryonic development. To date, as a potential drug target for discovering anti-tumor drugs, the medical significance of LSD1 has been greatly appreciated. However, the catalytic mechanism for the rate-limiting reductive half-reaction in demethylation remains controversial. By employing a combined computational approach including molecular modeling, molecular dynamics (MD) simulations and quantum mechanics/molecular mechanics (QM/MM) calculations, the catalytic mechanism of dimethylated H3K4 demethylation by LSD1 was characterized in details. The three-dimensional (3D) model of the complex was composed of LSD1, CoREST, and histone substrate. A 30-ns MD simulation of the model highlights the pivotal role of the conserved Tyr761 and lysine-water-flavin motif in properly orienting flavin adenine dinucleotide (FAD) with respect to substrate. The synergy of the two factors effectively stabilizes the catalytic environment and facilitated the demethylation reaction. On the basis of the reasonable consistence between simulation results and available mutagenesis data, QM/MM strategy was further employed to probe the catalytic mechanism of the reductive half-reaction in demethylation. The characteristics of the demethylation pathway determined by the potential energy surface and charge distribution analysis indicates that this reaction belongs to the direct hydride transfer mechanism. Our study provides insights into the LSD1 mechanism of reductive half-reaction in demethylation and has important implications for the discovery of regulators against LSD1 enzymes.
Conflict of interest statement
Figures


Similar articles
-
Lysine-specific histone demethylase 1 (LSD1): A potential molecular target for tumor therapy.Crit Rev Eukaryot Gene Expr. 2012;22(1):53-9. doi: 10.1615/critreveukargeneexpr.v22.i1.40. Crit Rev Eukaryot Gene Expr. 2012. PMID: 22339659 Review.
-
Amine oxidation mediated by lysine-specific demethylase 1: quantum mechanics/molecular mechanics insights into mechanism and role of lysine 661.J Am Chem Soc. 2013 Sep 11;135(36):13400-13. doi: 10.1021/ja403582u. Epub 2013 Aug 29. J Am Chem Soc. 2013. PMID: 23988016
-
Mining the Dynamical Properties of Substrate and FAD Binding Pockets of LSD1: Hints for New Inhibitor Design Direction.J Chem Inf Model. 2024 Jun 24;64(12):4773-4780. doi: 10.1021/acs.jcim.4c00398. Epub 2024 Jun 5. J Chem Inf Model. 2024. PMID: 38837697
-
Inhibition of Lysine-Specific Demethylase-1 (LSD1/KDM1A) Promotes the Adipogenic Differentiation of hESCs Through H3K4 Methylation.Stem Cell Rev Rep. 2016 Jun;12(3):298-304. doi: 10.1007/s12015-016-9650-z. Stem Cell Rev Rep. 2016. PMID: 27059868 Free PMC article.
-
[Histone demethylase LSD1 and its biological functions].Yi Chuan. 2010 Apr;32(4):331-8. doi: 10.3724/sp.j.1005.2010.00331. Yi Chuan. 2010. PMID: 20423886 Review. Chinese.
Cited by
-
Molecular dynamics simulations indicate an induced-fit mechanism for LSD1/CoREST-H3-histone molecular recognition.BMC Biophys. 2013 Nov 25;6(1):15. doi: 10.1186/2046-1682-6-15. BMC Biophys. 2013. PMID: 24274367 Free PMC article.
-
Inhibitors of Protein Methyltransferases and Demethylases.Chem Rev. 2018 Feb 14;118(3):989-1068. doi: 10.1021/acs.chemrev.6b00801. Epub 2017 Mar 24. Chem Rev. 2018. PMID: 28338320 Free PMC article. Review.
-
Theoretical insights into catalytic mechanism of protein arginine methyltransferase 1.PLoS One. 2013 Aug 20;8(8):e72424. doi: 10.1371/journal.pone.0072424. eCollection 2013. PLoS One. 2013. PMID: 23977297 Free PMC article.
-
[Antiproliferative effect of silencing LSD1 gene on Jurkat cell line and its mechanism].Zhonghua Xue Ye Xue Za Zhi. 2016 Jan;37(1):56-60. doi: 10.3760/cma.j.issn.0253-2727.2016.01.011. Zhonghua Xue Ye Xue Za Zhi. 2016. PMID: 26876255 Free PMC article. Chinese.
-
Exploration of 5-cyano-6-phenylpyrimidin derivatives containing an 1,2,3-triazole moiety as potent FAD-based LSD1 inhibitors.Acta Pharm Sin B. 2020 Sep;10(9):1658-1668. doi: 10.1016/j.apsb.2020.02.006. Epub 2020 Feb 24. Acta Pharm Sin B. 2020. PMID: 33088686 Free PMC article.
References
-
- Berger SL. The complex language of chromatin regulation during transcription. Nature. 2007;447:407–412. - PubMed
-
- Kouzarides T. Chromatin modifications and their function. Cell. 2007;128:693–705. - PubMed
-
- Shi Y, Lan F, Matson C, Mulligan P, Whetstine JR, et al. Histone demethylation mediated by the nuclear amine oxidase homolog LSD1. Cell. 2004;119:941–953. - PubMed
-
- Forneris F, Binda C, Vanoni MA, Mattevi A, Battaglioli E. Histone demethylation catalysed by LSD1 is a flavin-dependent oxidative process. FEBS Lett. 2005;579:2203–2207. - PubMed
-
- Huang J, Sengupta R, Espejo AB, Lee MG, Dorsey JA, et al. p53 is regulated by the lysine demethylase LSD1. Nature. 2007;449:105–108. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

