Characterization of the pleiotropic effects of the genotype G-specific 36-nucleotide insertion in the context of other hepatitis B virus genotypes
- PMID: 21994450
- PMCID: PMC3233120
- DOI: 10.1128/JVI.05583-11
Characterization of the pleiotropic effects of the genotype G-specific 36-nucleotide insertion in the context of other hepatitis B virus genotypes
Abstract
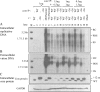
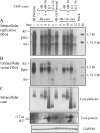
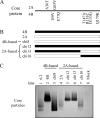
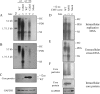
The pregenomic RNA (pgRNA) of hepatitis B virus (HBV) serves as the messenger for both core and P proteins, with the downstream P gene translated by ribosomal leaky scanning. HBV replication begins with packaging of the pgRNA and P protein into core protein particles, followed by conversion of RNA into DNA. Genotype G has a low replication capacity due to a low pgRNA level. It has a 36-nucleotide (nt) insertion in the 5' end of the core gene, adding 12 residues to the core protein. The insertion is needed to maintain efficient core protein expression and genome replication but causes inefficient virion secretion yet high maturity of virion DNA. In the present study, we confirmed that the 36-nt insertion had similar effects on core protein expression and virion secretion when it was introduced into genotype A and D clones but no impact on virion genome maturity. Surprisingly, the insertion impaired genome replication in both genotypes. Transcomplementation assays suggest that increased efficiency of core protein translation diminishes ribosomal scanning toward the downstream P gene. Indeed, mutating the core gene Kozak sequence restored core protein to lower levels but increased replication of the insertion mutant. Similar mutations impaired replication in genotype G. On the other hand, replacement of the core promoter sequence of genotype G with genotype A sequence increased pgRNA transcription and genome replication, implicating this region in the low replication capacity of genotype G. Why the 36-nt insertion is present in genotype G but absent in other genotypes is discussed.
Figures



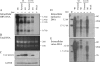
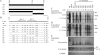
References
-
- Bang G., et al. 2005. Effect of mutating the two cysteines required for HBe antigenicity on hepatitis B virus DNA replication and virion secretion. Virology 332:216–224 - PubMed
-
- Bhat R. A., Ulrich P. P., Vyas G. N. 1990. Molecular characterization of a new variant of hepatitis B virus in a persistently infected homosexual man. Hepatology 11:271–276 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

