Functional characterization of alternative splicing in the C terminus of L-type CaV1.3 channels
- PMID: 21998309
- PMCID: PMC3234967
- DOI: 10.1074/jbc.M111.265207
Functional characterization of alternative splicing in the C terminus of L-type CaV1.3 channels
Abstract
Ca(V)1.3 channels are unique among the high voltage-activated Ca(2+) channel family because they activate at the most negative potentials and display very rapid calcium-dependent inactivation. Both properties are of crucial importance in neurons of the suprachiasmatic nucleus and substantia nigra, where the influx of Ca(2+) ions at subthreshold membrane voltages supports pacemaking function. Previously, alternative splicing in the Ca(V)1.3 C terminus gives rise to a long (Ca(V)1.3(42)) and a short form (Ca(V)1.3(42A)), resulting in a pronounced activation at more negative voltages and faster inactivation in the latter. It was further shown that the C-terminal modulator in the Ca(V)1.3(42) isoforms modulates calmodulin binding to the IQ domain. Using splice variant-specific antibodies, we determined that protein localization of both splice variants in different brain regions were similar. Using the transcript-scanning method, we further identified alternative splicing at four loci in the C terminus of Ca(V)1.3 channels. Alternative splicing of exon 41 removes the IQ motif, resulting in a truncated Ca(V)1.3 protein with diminished inactivation. Splicing of exon 43 causes a frameshift and exhibits a robust inactivation of similar intensity to Ca(V)1.3(42A). Alternative splicing of exons 44 and 48 are in-frame, altering interaction of the distal modulator with the IQ domain and tapering inactivation slightly. Thus, alternative splicing in the C terminus of Ca(V)1.3 channels modulates its electrophysiological properties, which could in turn alter neuronal firing properties and functions.
Figures
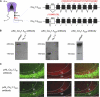




References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

