Radiogenomic mapping of edema/cellular invasion MRI-phenotypes in glioblastoma multiforme
- PMID: 21998659
- PMCID: PMC3187774
- DOI: 10.1371/journal.pone.0025451
Radiogenomic mapping of edema/cellular invasion MRI-phenotypes in glioblastoma multiforme
Erratum in
- PLoS One. 2012;7(2). doi:10.1371/annotation/b5267cb3-6aa7-47fc-a648-47f30a7cff3e. Majadan, Bhanu [corrected to Mahajan, Bhanu]
Abstract
Background: Despite recent discoveries of new molecular targets and pathways, the search for an effective therapy for Glioblastoma Multiforme (GBM) continues. A newly emerged field, radiogenomics, links gene expression profiles with MRI phenotypes. MRI-FLAIR is a noninvasive diagnostic modality and was previously found to correlate with cellular invasion in GBM. Thus, our radiogenomic screen has the potential to reveal novel molecular determinants of invasion. Here, we present the first comprehensive radiogenomic analysis using quantitative MRI volumetrics and large-scale gene- and microRNA expression profiling in GBM.
Methods: Based on The Cancer Genome Atlas (TCGA), discovery and validation sets with gene, microRNA, and quantitative MR-imaging data were created. Top concordant genes and microRNAs correlated with high FLAIR volumes from both sets were further characterized by Kaplan Meier survival statistics, microRNA-gene correlation analyses, and GBM molecular subtype-specific distribution.
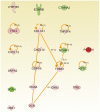
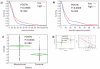
Results: The top upregulated gene in both the discovery (4 fold) and validation (11 fold) sets was PERIOSTIN (POSTN). The top downregulated microRNA in both sets was miR-219, which is predicted to bind to POSTN. Kaplan Meier analysis demonstrated that above median expression of POSTN resulted in significantly decreased survival and shorter time to disease progression (P<0.001). High POSTN and low miR-219 expression were significantly associated with the mesenchymal GBM subtype (P<0.0001).
Conclusion: Here, we propose a novel diagnostic method to screen for molecular cancer subtypes and genomic correlates of cellular invasion. Our findings also have potential therapeutic significance since successful molecular inhibition of invasion will improve therapy and patient survival in GBM.
Conflict of interest statement
Figures
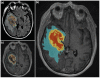
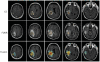
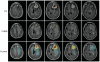


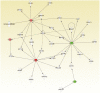
References
-
- Rickman DS, Bobek MP, Misek DE, Kuick R, Blaivas M, et al. Distinctive molecular profiles of high-grade and low-grade gliomas based on oligonucleotide microarray analysis. Cancer Res. 2001;61:6885–6891. - PubMed
-
- Mischel PS, Cloughesy TF, Nelson SF. DNA-microarray analysis of brain cancer: molecular classification for therapy. Nat Rev Neurosci. 2004;5:782–792. - PubMed
-
- Hegi ME, Diserens AC, Gorlia T, Hamou MF, de Tribolet N, et al. MGMT gene silencing and benefit from temozolomide in glioblastoma. N Engl J Med. 2005;352:997–1003. - PubMed
-
- Quant EC, Wen PY. Novel medical therapeutics in glioblastomas, including targeted molecular therapies, current and future clinical trials. Neuroimaging Clin N Am. 2010;20:425–448. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

