Defining the roles of cis-acting RNA elements in tombusvirus replicase assembly in vitro
- PMID: 22013057
- PMCID: PMC3255868
- DOI: 10.1128/JVI.00404-11
Defining the roles of cis-acting RNA elements in tombusvirus replicase assembly in vitro
Abstract
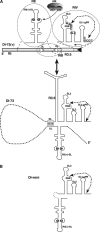
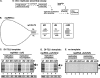
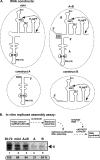
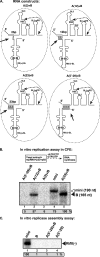
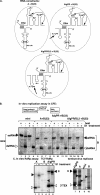
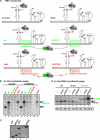
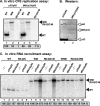
In addition to its central role as a template for replication and translation, the viral plus-strand RNA genome also has nontemplate functions, such as recruitment to the site of replication and assembly of the viral replicase, activities that are mediated by cis-acting RNA elements within viral genomes. Two noncontiguous RNA elements, RII(+)-SL (located internally in the tombusvirus genome) and RIV (located at the 3'-terminus), are involved in template recruitment into replication and replicase assembly; however, the importance of each of these RNA elements for these two distinct functions is not fully elucidated. We used an in vitro replicase assembly assay based on yeast cell extract and purified recombinant tombusvirus replication proteins to show that RII(+)-SL, in addition to its known requirement for recruitment of the plus-strand RNA into replication, is also necessary for assembly of an active viral replicase complex. Additional studies using a novel two-component RNA system revealed that the recruitment function of RII(+)-SL can be provided in trans by a separate RNA and that the replication silencer element, located within RIV, defines the template that is used for initiation of minus-strand synthesis. Collectively, this work has revealed new functions for tombusvirus cis-acting RNA elements and provided insights into the pioneering round of minus-strand synthesis.
Figures


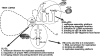
References
-
- An M, et al. 2010. A Y-shaped RNA structure in the 3′ untranslated region together with the trans-activator and core promoter of Red clover necrotic mosaic virus RNA2 is required for its negative-strand RNA synthesis. Virology 405:100–109 - PubMed
-
- Bertrand E, et al. 1998. Localization of ASH1 mRNA particles in living yeast. Mol. Cell 2:437–445 - PubMed
-
- den Boon JA, Ahlquist P. 2010. Organelle-like membrane compartmentalization of positive-strand RNA virus replication factories. Annu. Rev. Microbiol. 64:241–256 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

