Down-modulation of CD8αβ is a fundamental activity of primate lentiviral Nef proteins
- PMID: 22013062
- PMCID: PMC3255914
- DOI: 10.1128/JVI.00717-11
Down-modulation of CD8αβ is a fundamental activity of primate lentiviral Nef proteins
Abstract
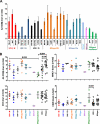
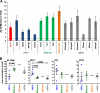
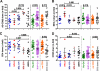
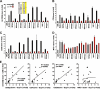
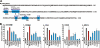
It is well established that the Nef proteins of human and simian immunodeficiency viruses (HIV and SIV) modulate major histocompatibility complex class I (MHC-I) cell surface expression to protect infected cells against lysis by cytotoxic T lymphocytes (CTLs). Recent data supported the observation that Nef also manipulates CTLs directly by down-modulating CD8αβ (J. A. Leonard, T. Filzen, C. C. Carter, M. Schaefer, and K. L. Collins, J. Virol. 85:6867-6881, 2011), but it remained unknown whether this Nef activity is conserved between different lineages of HIV and SIV. In this study, we examined a total of 42 nef alleles from 16 different primate lentiviruses representing most major lineages of primate lentiviruses, as well as nonpandemic HIV-1 strains and the direct precursors of HIV-1 (SIVcpz and SIVgor). We found that the vast majority of these nef alleles strongly down-modulate CD8β in human T cells. Primate lentiviral Nefs generally interacted specifically with the cytoplasmic tail of CD8β, and down-modulation of this receptor was dependent on the conserved dileucine-based motif and two adjacent acidic residues (DD/E) in the C-terminal flexible loop of SIV Nef proteins. Both of these motifs are known to be important for the interaction of HIV-1 Nef with AP-2, and they were also shown to be critical for down-modulation of CD4 and CD28, but not MHC-I, by SIV Nefs. Our results show that down-modulation of CD4, CD8β, and CD28 involves largely overlapping (but not identical) domains and is most likely dependent on conserved interactions of primate lentiviral Nefs with cellular adaptor proteins. Furthermore, our data demonstrate that Nef-mediated down-modulation of CD8αβ is a fundamental property of primate lentiviruses and suggest that direct manipulation of CD8+ T cells plays a relevant role in viral immune evasion.
Figures


Similar articles
-
Resistance of Major Histocompatibility Complex Class B (MHC-B) to Nef-Mediated Downregulation Relative to that of MHC-A Is Conserved among Primate Lentiviruses and Influences Antiviral T Cell Responses in HIV-1-Infected Individuals.J Virol. 2017 Dec 14;92(1):e01409-17. doi: 10.1128/JVI.01409-17. Print 2018 Jan 1. J Virol. 2017. PMID: 29046444 Free PMC article.
-
Down-modulation of primate lentiviral receptors by Nef proteins of simian immunodeficiency virus (SIV) of chimpanzees (SIVcpz) and related SIVs: implication for the evolutionary event at the emergence of SIVcpz.J Gen Virol. 2015 Sep;96(9):2867-2877. doi: 10.1099/vir.0.000207. Epub 2015 Jun 3. J Gen Virol. 2015. PMID: 26041873 Free PMC article.
-
Selective downmodulation of HLA-A and -B by Nef alleles from different groups of primate lentiviruses.Virology. 2008 Mar 30;373(1):229-37. doi: 10.1016/j.virol.2007.11.019. Epub 2007 Dec 21. Virology. 2008. PMID: 18155264
-
Role of Nef in primate lentiviral immunopathogenesis.Cell Mol Life Sci. 2008 Sep;65(17):2621-36. doi: 10.1007/s00018-008-8094-2. Cell Mol Life Sci. 2008. PMID: 18438604 Free PMC article. Review.
-
The downregulation of CD4 and MHC-I by primate lentiviruses: a paradigm for the modulation of cell surface receptors.Immunol Rev. 1999 Apr;168:51-63. doi: 10.1111/j.1600-065x.1999.tb01282.x. Immunol Rev. 1999. PMID: 10399064 Review.
Cited by
-
The HIV-1 accessory proteins Nef and Vpu downregulate total and cell surface CD28 in CD4+ T cells.Retrovirology. 2018 Jan 12;15(1):6. doi: 10.1186/s12977-018-0388-3. Retrovirology. 2018. PMID: 29329537 Free PMC article.
-
Tetherin downmodulation by SIVmac Nef lost with the H196Q escape variant is restored by an upstream variant.PLoS One. 2020 Aug 7;15(8):e0225420. doi: 10.1371/journal.pone.0225420. eCollection 2020. PLoS One. 2020. PMID: 32764749 Free PMC article.
-
How HIV Nef Proteins Hijack Membrane Traffic To Promote Infection.J Virol. 2019 Nov 26;93(24):e01322-19. doi: 10.1128/JVI.01322-19. Print 2019 Dec 15. J Virol. 2019. PMID: 31578291 Free PMC article. Review.
-
The KT Jeang Retrovirology prize 2016: Frank Kirchhoff.Retrovirology. 2016 Aug 5;13(1):53. doi: 10.1186/s12977-016-0286-5. Retrovirology. 2016. PMID: 27492506 Free PMC article. No abstract available.
-
Multifunctional Roles of the N-Terminal Region of HIV-1SF2Nef Are Mediated by Three Independent Protein Interaction Sites.J Virol. 2019 Dec 12;94(1):e01398-19. doi: 10.1128/JVI.01398-19. Print 2019 Dec 12. J Virol. 2019. PMID: 31597760 Free PMC article.
References
-
- Ariën KK, Verhasselt B. 2008. HIV Nef: role in pathogenesis and viral fitness. Curr. HIV Res. 6:200–208 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

