Analysis of the mating-type loci of co-occurring and phylogenetically related species of Ascochyta and Phoma
- PMID: 22014305
- PMCID: PMC6638728
- DOI: 10.1111/j.1364-3703.2011.00751.x
Analysis of the mating-type loci of co-occurring and phylogenetically related species of Ascochyta and Phoma
Abstract
Ascochyta and Phoma are fungal genera containing several important plant pathogenic species. These genera are morphologically similar, and recent molecular studies performed to unravel their phylogeny have resulted in the establishment of several new genera within the newly erected Didymellaceae family. An analysis of the structure of fungal mating-type genes can contribute to a better understanding of the taxonomic relationships of these plant pathogens, and may shed some light on their evolution and on differences in sexual strategy and pathogenicity. We analysed the mating-type loci of phylogenetically closely related Ascochyta and Phoma species (Phoma clematidina, Didymella vitalbina, Didymella clematidis, Peyronellaea pinodes and Peyronellaea pinodella) that co-occur on the same hosts, either on Clematis or Pisum. The results confirm that the mating-type genes provide the information to distinguish between the homothallic Pey. pinodes (formerly Ascochyta pinodes) and the heterothallic Pey. pinodella (formerly Phoma pinodella), and indicate the close phylogenetic relationship between these two species that are part of the disease complex responsible for Ascochyta blight on pea. Furthermore, our analysis of the mating-type genes of the fungal species responsible for causing wilt of Clematis sp. revealed that the heterothallic D. vitalbina (Phoma anamorph) is more closely related to the homothallic D. clematidis (Ascochyta anamorph) than to the heterothallic P. clematidina. Finally, our results indicate that homothallism in D. clematidis resulted from a single crossover between MAT1-1 and MAT1-2 sequences of heterothallic ancestors, whereas a single crossover event followed by an inversion of a fused MAT1/2 locus resulted in homothallism in Pey. pinodes.
© 2011 THE AUTHORS. MOLECULAR PLANT PATHOLOGY © 2011 BSPP AND BLACKWELL PUBLISHING LTD.
Figures

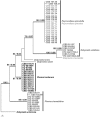
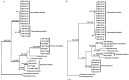


References
-
- Aveskamp, M.M. , De Gruyter, J. and Crous, P.W. (2008) Biology and recent developments in the systematics of Phoma, a complex genus of major quarantine significance. Fungal Divers. 31, 1–18.
-
- Aveskamp, M.M. , Verkley, G.J.M. , de Gruyter, J. , Murace, M.A. , Perello, A. , Woudenberg, J.H.C. , Groenewald, J.Z. and Crous, P.W. (2009a) DNA phylogeny reveals polyphyly of Phoma section Peyronellaea and multiple taxonomic novelties. Mycologia, 101, 363–382. - PubMed
-
- Aveskamp, M.M. , Woudenberg, J.H.C. , De Gruyter, J. , Turco, E. , Groenewald, J.Z. and Crous, P.W. (2009b) Development of taxon‐specific sequence characterized amplified region (SCAR) markers based on actin sequences and DNA amplification fingerprinting (DAF): a case study in the Phoma exigua species complex. Mol. Plant Pathol. 10, 403–414. - PMC - PubMed
Publication types
MeSH terms
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources

