Joint binding of OTX2 and MYC in promotor regions is associated with high gene expression in medulloblastoma
- PMID: 22016811
- PMCID: PMC3189962
- DOI: 10.1371/journal.pone.0026058
Joint binding of OTX2 and MYC in promotor regions is associated with high gene expression in medulloblastoma
Abstract
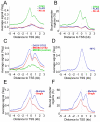
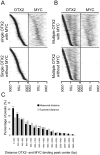
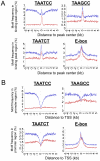
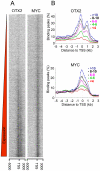
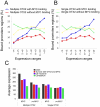
Both OTX2 and MYC are important oncogenes in medulloblastoma, the most common malignant brain tumor in childhood. Much is known about MYC binding to promoter regions, but OTX2 binding is hardly investigated. We used ChIP-on-chip data to analyze the binding patterns of both transcription factors in D425 medulloblastoma cells. When combining the data for all promoter regions in the genome, OTX2 binding showed a remarkable bi-modal distribution pattern with peaks around -250 bp upstream and +650 bp downstream of the transcription start sites (TSSs). Indeed, 40.2% of all OTX2-bound TSSs had more than one significant OTX2-binding peak. This OTX2-binding pattern was very different from the TSS-centered single peak binding pattern observed for MYC and other known transcription factors. However, in individual promoter regions, OTX2 and MYC have a strong tendency to bind in proximity of each other. OTX2-binding sequences are depleted near TSSs in the genome, providing an explanation for the observed bi-modal distribution of OTX2 binding. This contrasts to the enrichment of E-box sequences at TSSs. Both OTX2 and MYC binding independently correlated with higher gene expression. Interestingly, genes of promoter regions with multiple OTX2 binding as well as MYC binding showed the highest expression levels in D425 cells and in primary medulloblastomas. Genes within this class of promoter regions were enriched for medulloblastoma and stem cell specific genes. Our data suggest an important functional interaction between OTX2 and MYC in regulating gene expression in medulloblastoma.
Conflict of interest statement
Figures

References
-
- de Haas T, Oussoren E, Grajkowska W, Perek-Polnik M, Popovic M, et al. OTX1 and OTX2 expression correlates with the clinicopathologic classification of medulloblastomas. J Neuropathol Exp Neurol. 2006;65:176–186. - PubMed
-
- Michiels EM, Oussoren E, Van Groenigen M, Pauws E, Bossuyt PM, et al. Genes differentially expressed in medulloblastoma and fetal brain. Physiol Genomics. 1999;1:83–91. - PubMed
-
- Di C, Liao S, Adamson DC, Parrett TJ, Broderick DK, et al. Identification of OTX2 as a medulloblastoma oncogene whose product can be targeted by all-trans retinoic acid. Cancer Res. 2005;65:919–924. - PubMed
-
- Boon K, Eberhart CG, Riggins GJ. Genomic amplification of orthodenticle homologue 2 in medulloblastomas. Cancer Res. 2005;65:703–707. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

