Independently derived targeting of 28S rDNA by A- and D-clade R2 retrotransposons: Plasticity of integration mechanism
- PMID: 22016843
- PMCID: PMC3190273
- DOI: 10.4161/mge.1.1.16485
Independently derived targeting of 28S rDNA by A- and D-clade R2 retrotransposons: Plasticity of integration mechanism
Abstract
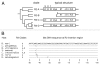
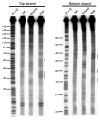
Restriction-like endonuclease (RLE) bearing non-LTR retrotransposons are site-specific elements that integrate into the genome through a target primed reverse transcription mechanism (TPRT). R2 elements have been used as a model system for investigating non-LTR retrotransposon integration. We previously demonstrated that R2 retrotransposons require two subunits of the element-encoded multifunctional protein to integrate-one subunit bound upstream of the insertion site and one bound downstream. R2 elements have been phylogenetically categorized into four clades: R2-A, B, C and D, that diverged from a common ancestor more than 850 million years ago. All R2 elements target the same sequence within 28S rDNA. The amino-terminal domain of R2Bm, an R2-D clade element, contains a single zinc finger and a Myb motif that are responsible for binding R2 protein downstream of the insertion site. Target site recognition is of interest as it is the first step in the integration reaction and may help elucidate evolutionary history and integration mechanism. The amino-terminal domain of R2-A clade members contains three zinc fingers and a Myb motif. We show here that R2Lp, an R2-A clade member, uses its amino-terminal DNA binding motifs to bind upstream of the insertion site. Because the R2-A and R2-D clade elements recognize 28S rDNA differently, we conclude the A- and D-clades represent independent targeting events to the 28S site. Our results also indicate a certain plasticity of insertional mechanics exists between the two clades.
Figures




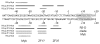
References
-
- Burke WD, Malik HS, Rich SM, Eickbush TH. Ancient lineages of non-LTR retrotransposons in the primitive eukaryote, Giardia lamblia. Mol Biol Evol. 2002;19:619–630. - PubMed
-
- Burke WD, Malik HS, Jones JP, Eickbush TH. The domain structure and retrotransposition mechanism of R2 elements are conserved throughout arthropods. Mol Biol Evol. 1999;16:502–511. - PubMed
-
- Kojima KK, Fujiwara H. Long-term inheritance of the 28S rDNA-specific retrotransposon R2. Mol Biol Evol. 2005;22:2157–2165. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
