Consensus among flexible fitting approaches improves the interpretation of cryo-EM data
- PMID: 22019767
- PMCID: PMC3288178
- DOI: 10.1016/j.jsb.2011.10.002
Consensus among flexible fitting approaches improves the interpretation of cryo-EM data
Abstract
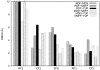
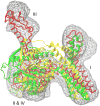
Cryo-elecron microscopy (cryo-EM) can provide important structural information of large macromolecular assemblies in different conformational states. Recent years have seen an increase in structures deposited in the Protein Data Bank (PDB) by fitting a high-resolution structure into its low-resolution cryo-EM map. A commonly used protocol for accommodating the conformational changes between the X-ray structure and the cryo-EM map is rigid body fitting of individual domains. With the emergence of different flexible fitting approaches, there is a need to compare and revise these different protocols for the fitting. We have applied three diverse automated flexible fitting approaches on a protein dataset for which rigid domain fitting (RDF) models have been deposited in the PDB. In general, a consensus is observed in the conformations, which indicates a convergence from these theoretically different approaches to the most probable solution corresponding to the cryo-EM map. However, the result shows that the convergence might not be observed for proteins with complex conformational changes or with missing densities in cryo-EM map. In contrast, RDF structures deposited in the PDB can represent conformations that not only differ from the consensus obtained by flexible fitting but also from X-ray crystallography. Thus, this study emphasizes that a "consensus" achieved by the use of several automated flexible fitting approaches can provide a higher level of confidence in the modeled configurations. Following this protocol not only increases the confidence level of fitting, but also highlights protein regions with uncertain fitting. Hence, this protocol can lead to better interpretation of cryo-EM data.
Copyright © 2011 Elsevier Inc. All rights reserved.
Figures

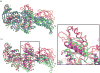
References
-
- Bahar I, Atilgan AR, Erman B. Direct evaluation of thermal fluctuations in proteins using a single-parameter harmonic potential. Fold Des. 1997;2:173–181. - PubMed
-
- Campbell ID. Timeline: the march of structural biology. Nat Rev Mol Cell Biol. 2002;3:377–381. - PubMed
-
- Chacon P, Wriggers W. Multi-resolution contour-based fitting of macromolecular structures. J Mol Biol. 2002;317:375–384. - PubMed
-
- Clementi C, Nymeyer H, Onuchic JN. Topological and energetic factors: what determines the structural details of the transition state ensemble and “en-route” intermediates for protein folding? An investigation for small globular proteins. J Mol Biol. 2000;298:937–953. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

