Endophytic life strategies decoded by genome and transcriptome analyses of the mutualistic root symbiont Piriformospora indica
- PMID: 22022265
- PMCID: PMC3192844
- DOI: 10.1371/journal.ppat.1002290
Endophytic life strategies decoded by genome and transcriptome analyses of the mutualistic root symbiont Piriformospora indica
Abstract
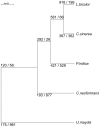
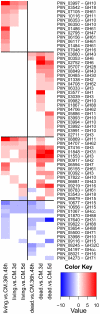
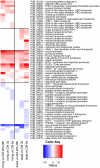
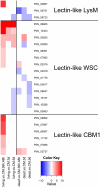
Recent sequencing projects have provided deep insight into fungal lifestyle-associated genomic adaptations. Here we report on the 25 Mb genome of the mutualistic root symbiont Piriformospora indica (Sebacinales, Basidiomycota) and provide a global characterization of fungal transcriptional responses associated with the colonization of living and dead barley roots. Extensive comparative analysis of the P. indica genome with other Basidiomycota and Ascomycota fungi that have diverse lifestyle strategies identified features typically associated with both, biotrophism and saprotrophism. The tightly controlled expression of the lifestyle-associated gene sets during the onset of the symbiosis, revealed by microarray analysis, argues for a biphasic root colonization strategy of P. indica. This is supported by a cytological study that shows an early biotrophic growth followed by a cell death-associated phase. About 10% of the fungal genes induced during the biotrophic colonization encoded putative small secreted proteins (SSP), including several lectin-like proteins and members of a P. indica-specific gene family (DELD) with a conserved novel seven-amino acids motif at the C-terminus. Similar to effectors found in other filamentous organisms, the occurrence of the DELDs correlated with the presence of transposable elements in gene-poor repeat-rich regions of the genome. This is the first in depth genomic study describing a mutualistic symbiont with a biphasic lifestyle. Our findings provide a significant advance in understanding development of biotrophic plant symbionts and suggest a series of incremental shifts along the continuum from saprotrophy towards biotrophy in the evolution of mycorrhizal association from decomposer fungi.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures











Similar articles
-
The root endophytic fungus Piriformospora indica requires host cell death for proliferation during mutualistic symbiosis with barley.Proc Natl Acad Sci U S A. 2006 Dec 5;103(49):18450-7. doi: 10.1073/pnas.0605697103. Epub 2006 Nov 20. Proc Natl Acad Sci U S A. 2006. PMID: 17116870 Free PMC article.
-
Host-related metabolic cues affect colonization strategies of a root endophyte.Proc Natl Acad Sci U S A. 2013 Aug 20;110(34):13965-70. doi: 10.1073/pnas.1301653110. Epub 2013 Aug 5. Proc Natl Acad Sci U S A. 2013. PMID: 23918389 Free PMC article.
-
A proteomics approach to study the molecular basis of enhanced salt tolerance in barley (Hordeum vulgare L.) conferred by the root mutualistic fungus Piriformospora indica.Mol Biosyst. 2013 Jun;9(6):1498-510. doi: 10.1039/c3mb70069k. Epub 2013 Apr 2. Mol Biosyst. 2013. PMID: 23545942
-
Opprimo ergo sum--evasion and suppression in the root endophytic fungus Piriformospora indica.Mol Plant Microbe Interact. 2012 Jun;25(6):727-37. doi: 10.1094/MPMI-11-11-0291. Mol Plant Microbe Interact. 2012. PMID: 22352718 Review.
-
Molecular mechanism underlying Piriformospora indica-mediated plant improvement/protection for sustainable agriculture.Acta Biochim Biophys Sin (Shanghai). 2019 Mar 1;51(3):229-242. doi: 10.1093/abbs/gmz004. Acta Biochim Biophys Sin (Shanghai). 2019. PMID: 30883651 Review.
Cited by
-
The plant strengthening root endophyte Piriformospora indica: potential application and the biology behind.Appl Microbiol Biotechnol. 2012 Dec;96(6):1455-64. doi: 10.1007/s00253-012-4506-1. Epub 2012 Oct 31. Appl Microbiol Biotechnol. 2012. PMID: 23108570 Free PMC article. Review.
-
Effector Profiles of Endophytic Fusarium Associated with Asymptomatic Banana (Musa sp.) Hosts.Int J Mol Sci. 2021 Mar 2;22(5):2508. doi: 10.3390/ijms22052508. Int J Mol Sci. 2021. PMID: 33801529 Free PMC article.
-
HSPRO controls early Nicotiana attenuata seedling growth during interaction with the fungus Piriformospora indica.Plant Physiol. 2012 Oct;160(2):929-43. doi: 10.1104/pp.112.203976. Epub 2012 Aug 14. Plant Physiol. 2012. PMID: 22892352 Free PMC article.
-
Functional Characterization of a Hexose Transporter from Root Endophyte Piriformospora indica.Front Microbiol. 2016 Jul 22;7:1083. doi: 10.3389/fmicb.2016.01083. eCollection 2016. Front Microbiol. 2016. PMID: 27499747 Free PMC article.
-
Structural plasticity in root-fungal symbioses: diverse interactions lead to improved plant fitness.PeerJ. 2018 Dec 4;6:e6030. doi: 10.7717/peerj.6030. eCollection 2018. PeerJ. 2018. PMID: 30533314 Free PMC article.
References
-
- Rodriguez RJ, White JF, Jr, Arnold AE, Redman RS. Fungal endophytes: diversity and functional roles. New Phytol. 2009;182:314–330. - PubMed
-
- Koide RT, Sharda JN, Herr JR, Malcolm GM. Ectomycorrhizal fungi and the biotrophy-saprotrophy continuum. New Phytol. 2008;178:230–233. - PubMed
-
- Schulz B, Boyle C. The endophytic continuum. Mycol Res. 2005;109:661–686. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

