Iron responsive mRNAs: a family of Fe2+ sensitive riboregulators
- PMID: 22026512
- PMCID: PMC3243817
- DOI: 10.1021/ar2001149
Iron responsive mRNAs: a family of Fe2+ sensitive riboregulators
Abstract
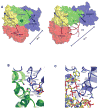
Messenger RNAs (mRNAs) are emerging as prime targets for small-molecule drugs. They afford an opportunity to assert control over an enormous range of biological processes: mRNAs regulate protein synthesis rates, have specific 3-D regulatory structures, and, in nucleated cells, are separated from DNA in space and time. All of the many steps between DNA copying (transcription) and ribosome binding (translation) represent potential control points. Messenger RNAs can fold into complex, 3-D shapes, such as tRNAs and rRNAs, providing an added dimension to the 2-D RNA structure (base pairing) targeted in many mRNA interference approaches. In this Account, we describe the structural and functional properties of the IRE (iron-responsive element) family, one of the few 3-D mRNA regulatory elements with known 3-D structure. This family of related base sequences regulates the mRNAs that encode proteins for iron metabolism. We begin by considering the IRE-RNA structure, which consists of a short (~30-nucleotide) RNA helix. Nature tuned the structure by combining a conserved AGU pseudotriloop, a closing C-G base pair, and a bulge C with various RNA helix base pairs. The result is a set of IRE-mRNAs with individual iron responses. The physiological iron signal is hexahydrated ferrous ion; in vivo iron responses vary over 10-fold depending on the individual IRE-RNA structure. We then discuss the interaction between the IRE-RNA structure and the proteins associated with it. IRE-RNA structures, which are usually noncoding, tightly bind specific proteins called IRPs. These repressor proteins are bound to IRE-RNA through C-bulge and AGU contacts that flip out a loop AG and a bulge C, bending the RNA helix. After binding, the exposed RNA surface then invites further interactions, such as with iron and other proteins. Binding of the IRE-RNA and the IRP also changes the IRP conformation. IRP binding stabilities vary 10-fold within the IRE family, reflecting individual IRE-RNA paired and unpaired bases. This variation contributes to the graded (hierarchical) iron responses in vivo. We also consider the mechanisms of IRE-mRNA control. The binding of Fe(2+) to IRE-RNA facilitates IRP release and the binding of eukaryotic initiation factors (eIFs), which are proteins that assemble mRNA, ribosomes, and tRNA for translation. IRE-RNAs are riboregulators for the inorganic metabolic signal, Fe(2+); they control protein synthesis rates by changing the distribution of the iron metabolic mRNAs between complexes with enhancing eIFs and inhibitory IRPs. The regulation of mRNA in the cytoplasm of eukaryotic cells is a burgeoning frontier in biomedicine. The evolutionarily refined IRE-RNAs, although absent in plants and bacteria, constitute a model system for 3-D mRNAs in all organisms. IRE-mRNAs have yielded "proof of principle" data for small-molecule targeting of mRNA structures, demonstrating tremendous potential for chemical manipulation of mRNA and protein synthesis in living systems.
Figures


Similar articles
-
IRE mRNA riboregulators use metabolic iron (Fe(2+)) to control mRNA activity and iron chemistry in animals.Metallomics. 2015 Jan;7(1):15-24. doi: 10.1039/c4mt00136b. Epub 2014 Sep 11. Metallomics. 2015. PMID: 25209685 Review.
-
Fe2+ binds iron responsive element-RNA, selectively changing protein-binding affinities and regulating mRNA repression and activation.Proc Natl Acad Sci U S A. 2012 May 29;109(22):8417-22. doi: 10.1073/pnas.1120045109. Epub 2012 May 14. Proc Natl Acad Sci U S A. 2012. PMID: 22586079 Free PMC article.
-
Loops and bulge/loops in iron-responsive element isoforms influence iron regulatory protein binding. Fine-tuning of mRNA regulation?J Biol Chem. 1998 Sep 11;273(37):23637-40. doi: 10.1074/jbc.273.37.23637. J Biol Chem. 1998. PMID: 9726965
-
Internal loop/bulge and hairpin loop of the iron-responsive element of ferritin mRNA contribute to maximal iron regulatory protein 2 binding and translational regulation in the iso-iron-responsive element/iso-iron regulatory protein family.Biochemistry. 2000 May 23;39(20):6235-42. doi: 10.1021/bi9924765. Biochemistry. 2000. PMID: 10821699
-
The IRE (iron regulatory element) family: structures which regulate mRNA translation or stability.Biofactors. 1993 May;4(2):87-93. Biofactors. 1993. PMID: 8347279 Review.
Cited by
-
RNA folding and catalysis mediated by iron (II).PLoS One. 2012;7(5):e38024. doi: 10.1371/journal.pone.0038024. Epub 2012 May 31. PLoS One. 2012. PMID: 22701543 Free PMC article.
-
Ferritin protein nanocage ion channels: gating by N-terminal extensions.J Biol Chem. 2012 Apr 13;287(16):13016-25. doi: 10.1074/jbc.M111.332734. Epub 2012 Feb 23. J Biol Chem. 2012. PMID: 22362775 Free PMC article.
-
Before It Gets Started: Regulating Translation at the 5' UTR.Comp Funct Genomics. 2012;2012:475731. doi: 10.1155/2012/475731. Epub 2012 May 28. Comp Funct Genomics. 2012. PMID: 22693426 Free PMC article.
-
A theranostic abscisic acid-based molecular glue.Chem Sci. 2023 Feb 28;14(12):3377-3384. doi: 10.1039/d2sc06995d. eCollection 2023 Mar 22. Chem Sci. 2023. PMID: 36970087 Free PMC article.
-
Gram-Negative Bacterial Envelope Homeostasis under Oxidative and Nitrosative Stress.Microorganisms. 2022 Apr 28;10(5):924. doi: 10.3390/microorganisms10050924. Microorganisms. 2022. PMID: 35630368 Free PMC article. Review.
References
-
- Bocobza S, Aharoni A. Switching the light on plant riboswitches. Trends Plant Sci. 2008;13:526–533. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

